Overcoming Biofouling and Enhancing Sensor Stability: The Path to Reliable Next-Generation Wearable Biosensors
This article provides a comprehensive analysis for researchers and drug development professionals on the critical challenges of biofouling and signal instability in wearable biosensors.
Overcoming Biofouling and Enhancing Sensor Stability: The Path to Reliable Next-Generation Wearable Biosensors
Abstract
This article provides a comprehensive analysis for researchers and drug development professionals on the critical challenges of biofouling and signal instability in wearable biosensors. We explore the fundamental mechanisms behind sensor performance degradation, review cutting-edge methodological approaches for surface engineering and antifouling strategies, detail practical troubleshooting and optimization protocols, and evaluate validation frameworks for comparing sensor performance in complex biological matrices. The aim is to synthesize current research into a practical guide for developing robust, long-term monitoring devices for biomedical research and clinical applications.
Understanding the Enemy: The Fundamental Science of Biofouling and Sensor Degradation
Technical Support & Troubleshooting Center
Frequently Asked Questions (FAQ)
Q1: My wearable biosensor's sensitivity decreases by over 50% within 6 hours of continuous skin contact. Is this biofouling or intrinsic sensor instability? A: This rapid decline is highly indicative of early-stage biofouling. Proteins (e.g., albumin, fibrinogen) and lipids from sweat and interstitial fluid form an occlusive layer on the sensing interface, physically blocking analyte access. Intrinsic instability (e.g., enzyme degradation, electrode passivation) typically manifests as a slower, more linear drift (>24-48 hrs). Perform a control experiment with a fouling-mimetic solution (see Protocol 1) to isolate the variable.
Q2: What are the primary chemical contributors to biofouling in sweat-sensing devices? A: Based on recent proteomic and metabolomic analyses of the skin-sensor interface, the key contributors are:
| Contributor Category | Specific Examples | Typical Concentration Range (in Sebum/Sweat) | Primary Impact on Sensor |
|---|---|---|---|
| Proteins | Albumin, Immunoglobulin G, Lysozyme, Collagen fragments | 10-500 µg/mL | Forms an adhesive, insulating layer; non-specific binding. |
| Lipids | Squalene, Triglycerides, Free Fatty Acids (e.g., Oleic acid) | 100-1000 µg/cm² (skin surface) | Hydrophobic layer causes signal drift; fouls hydrophobic membranes. |
| Cells & Debris | Corneocytes (skin cells), Microbes (S. epidermidis, C. acnes) | Variable | Physical barrier; microbial metabolism alters local pH/analyte concentration. |
| Electrolytes | Na⁺, K⁺, Cl⁻, Ca²⁺ | 10-100 mM (sweat) | Can cause crystallization on electrodes; alter electrochemical baseline. |
Q3: How can I distinguish signal drift due to sensor material degradation from drift due to biofouling? A: Implement a standardized differential measurement protocol. Use a two-electrode system: one functionalized sensing electrode and one identical "sentinel" electrode passivated to be non-responsive to the target analyte but exposed to the same biofouling environment. The drift in the sentinel electrode's non-faradaic impedance or baseline current is primarily due to biofouling. Subtract this from the sensing electrode's total drift to estimate material-based instability.
Q4: My anti-fouling hydrogel coating successfully reduces protein adsorption but drastically increases the sensor's response time. How can I mitigate this? A: This is a classic trade-off. The increased diffusional barrier of the hydrogel is slowing analyte transport. Consider these solutions:
- Optimize Crosslink Density: Reduce the hydrogel's crosslinking percentage to increase mesh size, facilitating faster diffusion.
- Layer Thickness: Ensure your coating is < 5 µm. Use spin-coating or electrodeposition for thin, uniform layers.
- Hydrophilic-Hydrophobic Balance: Incorporate a small fraction of hydrophobic monomers to prevent excessive swelling, which elongates diffusion paths.
Experimental Protocols
Protocol 1: In Vitro Fouling-Mimetic Challenge Test
Purpose: To accelerate and standardize the testing of a wearable sensor's susceptibility to biofouling. Materials: Biosensor prototype, potentiostat, artificial sweat (ISO 3160-2 standard), 2 g/L BSA (Bovine Serum Albumin) + 0.2 g/L Lysozyme in PBS, 0.1 mM Squalene in ethanol. Procedure:
- Baseline Measurement: Record the sensor's stable baseline and calibrated response to a target analyte (e.g., 0.1 mM glucose, 10 µM cortisol) in clean PBS.
- Fouling Layer Formation: Immerse the sensor in the BSA/Lysozyme solution for 30 minutes at 37°C. Rinse gently with PBS.
- Lipid Deposition: Gently pipette 50 µL of the Squalene solution onto the sensor surface and allow to air dry for 5 minutes.
- Post-Fouling Measurement: Immediately repeat the analyte response measurement from Step 1 using the same concentrations.
- Quantification: Calculate the percentage reduction in sensitivity (ΔI/ΔC) and increase in response time (T90).
Protocol 2: Real-Time Impedance Monitoring for Instability Diagnosis
Purpose: To non-destructively monitor the onset of biofouling and material degradation in situ. Materials: Biosensor with integrated interdigitated electrodes (IDEs), electrochemical impedance spectrometer (EIS). Procedure:
- Pre-Deployment Scan: Perform an EIS scan (e.g., 100 kHz to 1 Hz, 10 mV amplitude) on the clean, dry sensor in air. Record the baseline impedance magnitude at a high frequency (R∞) and low frequency (Rs).
- In-Use Monitoring: During wear or in vitro testing, perform brief EIS scans at regular intervals (e.g., every 15 minutes). Focus on a single low frequency (e.g., 1 Hz).
- Data Interpretation:
- A steady increase in low-frequency impedance is strongly correlated with the formation of an insulating biofouling layer.
- A steady increase in high-frequency impedance often indicates cracking, delamination, or bulk degradation of the sensor's conductive materials.
- Plot normalized impedance versus time to create a stability fingerprint.
Visualizations
Diagnostic Decision Tree for Signal Drift
Experimental Workflow for Coating Validation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Primary Function | Example Use-Case & Rationale |
|---|---|---|
| Poly(ethylene glycol) (PEG)-based Thiols (e.g., SH-PEG-SH) | Forms anti-fouling self-assembled monolayers (SAMs) on gold electrodes. Creates a hydrophilic, neutrally charged barrier to protein adsorption. | Gold electrode functionalization. The thiol binds to Au, while the dense PEG brush sterically repels biomolecules. |
| Zwitterionic Polymers (e.g., Poly(sulfobetaine methacrylate) (pSBMA)) | Ultra-hydrophilic, electrostatically neutral coating that binds water molecules tightly via ionic solvation, preventing foulant adhesion. | Hydrogel or polymer brush coating for long-term wearable patches. Superior long-term stability vs. PEG in wet environments. |
| Artificial Eccrine Sweat (ISO 3160-2) | Standardized electrolyte solution for baseline performance and stability testing. Contains NaCl, urea, lactate, etc. | In vitro calibration and control experiments to isolate effects of electrolytes from organic foulants. |
| Quartz Crystal Microbalance with Dissipation (QCM-D) | Label-free, real-time measurement of mass (ng/cm²) and viscoelastic properties of adlayers on sensor surfaces. | Quantifying the kinetics and mass of protein adsorption (e.g., BSA, fibrinogen) onto novel coating materials. |
| Faradaic EIS Redox Probes (e.g., [Fe(CN)₆]³⁻/⁴⁻) | Electroactive probe to monitor changes in electron transfer kinetics at the electrode-coating interface. | Diagnosing coating failure. An increase in charge-transfer resistance (Rct) indicates fouling or coating degradation blocking the probe. |
| Skin-like Substrate (e.g., PDMS with Micropillars) | Elastomeric substrate mimicking skin's modulus, topography, and sweat transport. | Testing mechanical stability (cracking/delamination) and sweat wicking in a realistic form factor during benchtop tests. |
Technical Support Center: Troubleshooting for Biofouling & Sensor Stability Research
Frequently Asked Questions (FAQs)
Q1: Our QCM-D (Quartz Crystal Microbalance with Dissipation) data shows inconsistent frequency (ΔF) and dissipation (ΔD) shifts during initial protein adsorption experiments. What could be causing this? A: Inconsistent ΔF/ΔD shifts often indicate problems with surface preparation or solution conditions.
- Cause 1: Inadequate or inconsistent sensor surface cleaning prior to coating. Residual contaminants compete for binding.
- Solution: Implement a strict UV-ozone or oxygen plasma cleaning protocol for 10-15 minutes immediately before use. Follow with precise buffer rinsing.
- Cause 2: Uncontrolled flow rate or air bubbles in the fluidic system.
- Solution: Use a syringe pump with calibrated flow rates (typically 50-100 µL/min). Degas all buffers and protein solutions before injection. Include a bubble trap in-line.
- Cause 3: Protein solution aggregation or denaturation.
- Solution: Centrifuge protein aliquots at 14,000 x g for 10 minutes at 4°C before use. Avoid repeated freeze-thaw cycles.
Q2: Our anti-fouling polymer brush coatings (e.g., PEG, zwitterions) show great performance in lab buffer but fail rapidly in complex biofluids (e.g., serum, sweat). How can we improve stability? A: Failure in complex media is often due to coating degradation or "stealth" failure.
- Cause 1: Physiologically relevant ionic strengths can collapse or alter polymer brush conformation, reducing steric repulsion.
- Solution: Optimize brush density and chain length. Consider mixed-charge zwitterionic polymers (e.g., poly(carboxybetaine)) which maintain hydration across a wider ionic range.
- Cause 2: Enzymatic or oxidative degradation of the coating in biofluids.
- Solution: Use more robust backbone chemistries (e.g., peptidomimetics) or incorporate cross-linking within the brush layer. Test coating stability in target biofluid in vitro prior to sensor integration.
Q3: During live-cell adhesion assays, we observe high variability in adherent cell counts between identical antifouling test surfaces. What are the key controls? A: Variability typically stems from cell handling or surface conditioning.
- Cause 1: Inconsistent cell seeding density or viability.
- Solution: Always use a hemocytometer or automated cell counter to standardize seeding density. Confirm viability >95% with Trypan Blue exclusion. Use cells at low passage number.
- Cause 2: Uncontrolled formation of a "conditioning film" of adsorbed proteins from the cell culture medium prior to cell contact.
- Solution: Pre-incubate all test surfaces in the specific serum-containing medium for a fixed time (e.g., 30 min) before adding cells. This standardizes the starting point for cell adhesion.
Q4: Our electrochemical biosensor signals drift rapidly upon exposure to biological samples, complicating data interpretation. How can we distinguish biofouling from other signal loss? A: Implement control experiments to deconvolute signal loss mechanisms.
- Solution Protocol:
- Fouling Control: Use a non-functionalized version of your sensor (blocked active site) alongside the active sensor. Signal change on the control is primarily due to biofouling (non-specific).
- Stability Control: Run the active sensor in a sterile, protein-free buffer (e.g., PBS). Any signal drift is due to sensor instability (e.g., reference electrode drift, enzyme leaching).
- Specificity Control: Test the sensor in the sample after specifically removing the target analyte (e.g., via filtration, digestion). Remaining signal is interference.
Experimental Protocols
Protocol 1: Standardized QCM-D Assay for Protein Adsorption Kinetics
- Objective: Quantify the mass and viscoelastic properties of an adsorbed protein layer on a test coating.
- Materials: QCM-D instrument (e.g., Biolin Scientific), gold-coated quartz sensors, UV-ozone cleaner, phosphate-buffered saline (PBS, pH 7.4), protein solution (e.g., fibrinogen, 1 mg/mL in PBS).
- Method:
- Clean sensor chips in UV-ozone for 15 minutes.
- Mount chip, initiate flow of PBS at 100 µL/min until stable baseline (ΔF < 0.5 Hz over 10 min).
- Switch flow to protein solution for 30 minutes.
- Switch back to PBS flow for 15 minutes to rinse off loosely bound protein.
- Record ΔF (frequency shift, related to mass) and ΔD (dissipation shift, related to rigidity) at the 3rd, 5th, and 7th overtones. Data from the 7th overtone is typically used for analysis.
Protocol 2: Static Bacterial Adhesion Assay for Antifouling Surfaces
- Objective: Quantify the number of adherent bacterial cells after a fixed incubation period.
- Materials: Test substrates (e.g., coated sensor surfaces), bacterial culture (e.g., P. aeruginosa PAO1), tryptic soy broth (TSB), sterile PBS, crystal violet stain (0.1% w/v), acetic acid (33% v/v), 24-well plate.
- Method:
- Grow bacteria overnight in TSB, dilute 1:100 in fresh TSB, and grow to mid-log phase (OD600 ≈ 0.5).
- Wash bacterial cells twice in PBS and resuspend in TSB or minimal medium to ~1 x 10^7 CFU/mL.
- Place test substrates in wells of a 24-well plate. Add 2 mL of bacterial suspension per well. Incubate statically at 37°C for 2 hours.
- Gently rinse each substrate 3 times with PBS to remove non-adherent cells.
- Fix adherent cells with 2 mL methanol for 15 minutes. Air dry.
- Stain with 1 mL crystal violet for 5 minutes. Rinse thoroughly with water.
- Destain with 2 mL of 33% acetic acid for 15 minutes with gentle shaking.
- Measure OD590 of the destained solution. Correlate to a standard curve of known bacterial counts.
Protocol 3: Electrochemical Impedance Spectroscopy (EIS) for Monitoring Biofilm Formation
- Objective: Monitor the progression of biofilm growth on an electrode surface in real-time.
- Materials: Potentiostat with EIS capability, 3-electrode system (working electrode = test substrate, Pt counter electrode, Ag/AgCl reference), growth medium.
- Method:
- Set up the electrochemical cell with the test substrate as the working electrode in sterile growth medium.
- Record a baseline EIS spectrum (e.g., 10^5 Hz to 0.1 Hz, 10 mV RMS amplitude) at open circuit potential.
- Inoculate the medium with the target microorganism (e.g., S. epidermidis).
- At regular intervals (e.g., every 30 minutes for 24-48 hours), pause agitation and record a new EIS spectrum.
- Fit spectra to a modified Randles equivalent circuit. The increase in charge transfer resistance (Rct) is a direct quantitative measure of biofilm insulation on the electrode surface.
Data Presentation
Table 1: Efficacy of Common Antifouling Coatings in Model Systems
| Coating Type | Example Material | ΔF on QCM-D (Fibrinogen) | Bacterial Adhesion Reduction vs. Bare Au | Stability in Serum (Days) | Key Limitation |
|---|---|---|---|---|---|
| Polyethylene Glycol | PEG-Thiol (5k Da) | -25 ± 3 Hz | 85 ± 5% | 1-2 | Oxidative degradation |
| Zwitterionic Polymer | Poly(SBMA) brush | -8 ± 2 Hz | 95 ± 3% | 7-10 | Sensitive to pH extremes |
| Hydrophilic Peptide | EKEKEKE-PEP | -15 ± 4 Hz | 80 ± 7% | 3-5 | Proteolytic cleavage |
| Antifouling Hydrogel | PHEMA-based | -5 ± 1 Hz* | 90 ± 4% | 14+ | Can slow analyte diffusion |
Note: Data are representative values from recent literature (2023-2024). ΔF measured at 5th overtone. Stability defined as <10% loss of antifouling performance.
Table 2: Impact of Biofouling Cascade on Electrochemical Biosensor Performance
| Fouling Stage | Sensor Parameter Affected | Typical Signal Drift (in 10% Serum) | Reversibility |
|---|---|---|---|
| Protein Adsorption (Minutes) | Baseline Current/Noise | +5 to 15% | Irreversible |
| Cell Adhesion (Hours) | Sensitivity (Slope) | -20 to 40% | Partially Reversible |
| Biofilm Formation (Days) | Charge Transfer Resistance (Rct) | +200 to 1000% | Irreversible |
Visualizations
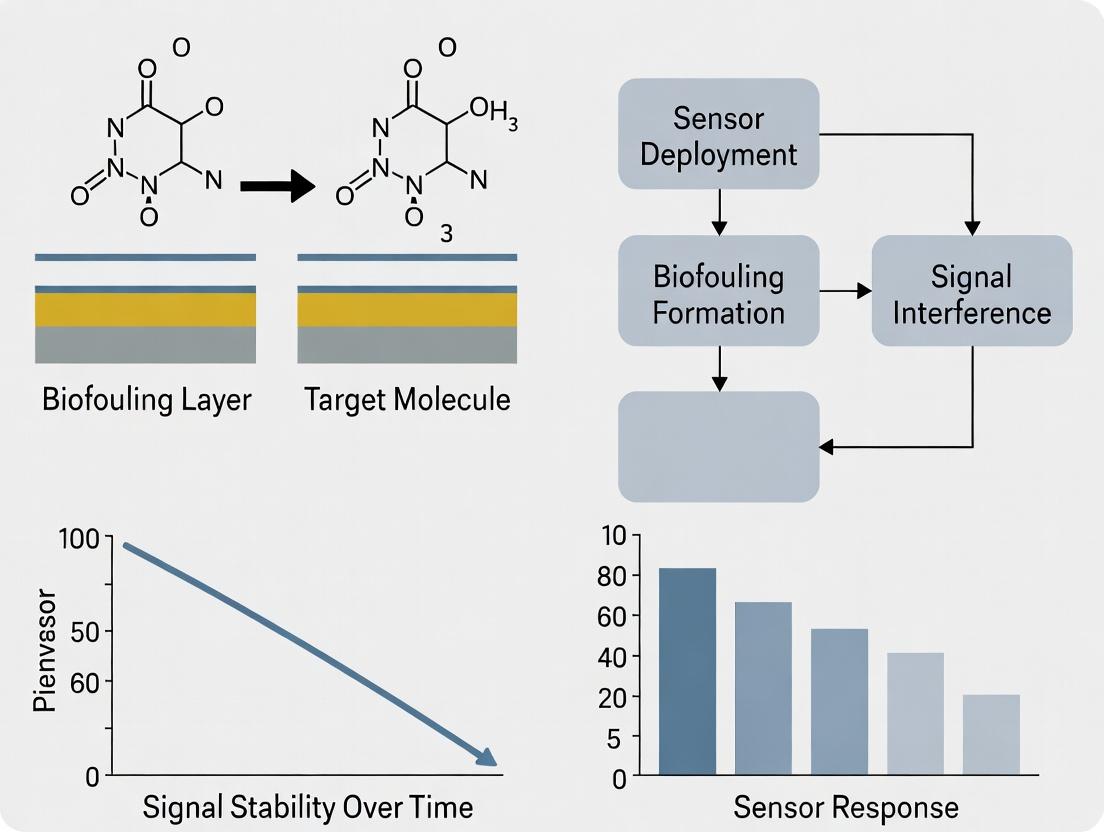
Diagram 1: Biofouling Cascade Impact on Sensor
Diagram 2: Experimental Workflow for Fouling Analysis
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Biofouling Research | Example Product/Chemical |
|---|---|---|
| Gold-coated QCM-D Sensors | Standardized substrate for protein adsorption kinetics studies. Can be functionalized with various coatings. | Biolin Scientific QSX 301 Gold sensors. |
| Zwitterionic Monomer (SBMA) | Synthesis of ultra-low fouling polymer brush coatings via surface-initiated ATRP. | [2-(Methacryloyloxy)ethyl]dimethyl-(3-sulfopropyl)ammonium hydroxide (SBMA). |
| Pluronic F-127 | Non-ionic surfactant used for blocking non-specific adsorption and as a temporary antifouling layer. | Often used in 0.1-1% w/v solution to passivate surfaces and microfluidic channels. |
| Crystal Violet Stain | Quantitative and qualitative analysis of adherent bacterial or fungal biomass on surfaces. | 0.1% aqueous crystal violet solution for staining fixed biofilms. |
| Fibronectin, Fibrinogen | Model "sticky" proteins for challenging antifouling surfaces in adsorption experiments. | Human plasma-derived proteins, used at 1 mg/mL in PBS. |
| Polyethylene Glycol Thiol (PEG-SH) | Gold-standard for creating protein-resistant monolayers on gold surfaces via self-assembly. | HS-C11-EG6-OH (e.g., from Sigma-Aldrich or Nanocs). |
| DAPI Stain | Fluorescent nuclear stain for quantifying adherent mammalian cell numbers on test substrates. | 4',6-diamidino-2-phenylindole, used at 1 µg/mL for microscopy. |
| Electrochemical Redox Probe | For EIS and voltammetry to monitor biofilm-induced insulation. | [Fe(CN)6]3−/4− in PBS, typically 5 mM each. |
Troubleshooting Guides & FAQs
Q1: How can I differentiate between signal drift caused by electrode passivation versus enzyme denaturation? A1: Perform a two-step diagnostic protocol. First, run a standard ferri/ferrocyanide redox probe test. A >20% increase in peak-to-peak separation in cyclic voltammetry indicates significant electrode passivation. Second, after recalibrating the electrode in fresh buffer, add a known concentration of substrate. A recovery of <80% of the expected current suggests concurrent enzyme denaturation. The table below summarizes the diagnostic signatures:
| Observed Signal Change | Redox Probe Test Result | Post-Buffer Calibration Response | Likely Primary Cause |
|---|---|---|---|
| Gradual, monotonic decrease | Normal | Normal | Enzyme Denaturation |
| Gradual, noisy decrease | Increased Peak Separation | Low | Electrode Passivation |
| Sharp initial drop, then gradual | Increased Peak Separation | Very Low | Combined Passivation & Denaturation |
Experimental Protocol: Redox Probe Diagnostic
- Reagents: 5 mM Potassium ferricyanide, 5 mM Potassium ferrocyanide, 0.1 M KCl supporting electrolyte.
- Procedure: Record a cyclic voltammogram (CV) from -0.1 V to +0.5 V vs. Ag/AgCl at 50 mV/s in the redox solution.
- Measurement: Calculate the peak-to-peak separation (ΔEp). A fresh, clean electrode shows ΔEp ~59-70 mV. A value >85 mV indicates fouling/passivation.
Q2: What are effective in-situ methods to mitigate membrane fouling in continuous wearable biosensors? A2: Current research focuses on surface modifications and electrochemical cleaning cycles. Implementing a low-frequency (e.g., 0.1 Hz) -0.4 V vs. Ag/AgCl pulse for 30 seconds every 30 minutes can reduce protein adsorption by ~40% in vitro. Furthermore, coating the outer membrane with a hydrogel layer containing polyethylene glycol (PEG) can reduce biofouling from complex fluids (e.g., artificial sweat) by 60-70% over 24 hours compared to uncoated surfaces.
Experimental Protocol: Anti-Fouling Hydrogel Coating
- Reagents: Polyethylene glycol diacrylate (PEGDA, 700 Da), Photoinitiator (Irgacure 2959), Phosphate Buffered Saline (PBS).
- Procedure: Mix PEGDA (20% w/v) with photoinitiator (0.5% w/v) in PBS. Dip-coat the sensor membrane and cure under UV light (365 nm, 5 mW/cm²) for 3 minutes.
- Validation: Soak coated and uncoated sensors in fluorescein-labeled bovine serum albumin (BSA) solution (1 mg/mL) for 2 hours. Measure fluorescence intensity adhered to the membrane; a >60% reduction confirms coating efficacy.
Q3: What are the quantitative indicators of enzyme denaturation in a biosensor layer? A3: Monitor the change in apparent Michaelis-Menten constant (Km) and maximum current (Imax). Denaturation typically causes a decrease in Imax due to loss of active enzyme, while Km may increase due to hindered substrate diffusion through a degrading matrix. A 30% reduction in Imax over 8 hours of continuous operation at 37°C is a common benchmark for instability.
| Parameter | Fresh Sensor | After 8-hr Operation | % Change | Interpretation |
|---|---|---|---|---|
| Imax (nA) | 100 ± 5 | 68 ± 7 | -32% | Significant enzyme loss |
| Km (mM) | 2.0 ± 0.2 | 3.1 ± 0.3 | +55% | Increased diffusion barrier |
| Response Time (s) | 15 ± 2 | 25 ± 4 | +67% | Matrix degradation |
Experimental Protocol: Kinetic Parameter Estimation
- Procedure: Measure steady-state current response in standard buffer with increasing substrate concentrations (e.g., 0.1, 0.5, 1, 2, 5, 10 mM).
- Analysis: Fit data to the Michaelis-Menten equation: I = (I_max * [S]) / (K_m + [S]) using non-linear regression software.
- Tracking: Perform this calibration at time zero and after defined operational periods.
Diagnostic Workflow for Signal Drift
Anti-Fouling Mitigation Strategies
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Potassium Ferri/Ferrocyanide | Redox probe for diagnosing electron transfer kinetics at the electrode surface. An increased peak separation indicates passivation. |
| Polyethylene Glycol Diacrylate (PEGDA) | Hydrogel precursor for creating anti-fouling, hydrophilic coatings that reduce non-specific protein adsorption on sensor membranes. |
| Irgacure 2959 | A biocompatible photoinitiator used to crosslink PEGDA hydrogel coatings upon exposure to UV light. |
| Fluorescein-Labeled BSA | Model foulant protein. Its fluorescence allows for quantitative measurement of protein adsorption on sensor surfaces. |
| Appropriate Enzyme Substrate | Used in kinetic studies (Imax, Km) to quantify the extent of enzyme denaturation within the biosensor layer. |
| Artificial Interstitial Fluid/Sweat | Complex, ion-containing fluid for in vitro testing of sensor stability and fouling under physiologically relevant conditions. |
Impact on Pharmacokinetic/Pharmacodynamic (PK/PD) Studies and Clinical Data Integrity
Technical Support Center
Frequently Asked Questions (FAQs) & Troubleshooting
Q1: Our wearable biosensor shows signal drift during long-term (>24h) continuous monitoring, potentially corrupting PK profiles. What are the primary causes and immediate corrective actions? A: Signal drift is frequently caused by progressive biofouling (protein adsorption, cellular adhesion) and reference electrode instability. Immediate actions include:
- Recalibration: Perform a two-point in-situ calibration using fresh analyte standards if the device platform allows it.
- Data Post-Processing: Apply baseline correction algorithms (e.g., moving average, high-pass filter) to the raw signal, noting this as a limitation in your methods.
- Inspect Physically: Check for visible biofilm formation or delamination of the sensor membrane. The experiment may need to be terminated.
Q2: How does biofouling specifically impact the pharmacodynamic (PD) endpoints derived from interstitial fluid (ISF) measurements? A: Biofouling creates a diffusion barrier between the ISF and the sensor's recognition element. This leads to:
- Increased Lag Time: A delayed sensor response relative to the actual blood/ISF concentration change, distorting the
T_maxand kinetic shape of the PD response curve. - Signal Attenuation: Reduced amplitude of the measured signal, leading to an underestimation of
C_maxor effect magnitude. - Corrective Protocol: Conduct a pilot study to characterize the time-dependent lag by comparing sensor readings with frequent, paired microdialysis or blood samples to establish a correction model.
Q3: What experimental protocols can we implement during study design to proactively monitor and account for sensor performance decay? A: Implement a dual-validation protocol:
- Internal QC Spikes: If the sensor design permits, introduce a known concentration of a control analyte (non-pharmacological) at regular intervals (e.g., every 12 hours via integrated microneedles) to measure recovery rate.
- Paired Sampling: Schedule periodic gold-standard blood draws (e.g., at trough, peak, and once during elimination phase) to correlate and calibrate the continuous sensor data.
- Pre- and Post-Study Calibration: Characterize each sensor's response in vitro before deployment and after explanation to quantify signal loss.
Q4: We suspect inflammation from the wearable device is altering local tissue permeability and thus PK/PD readings. How can we troubleshoot this? A: Local inflammation can change capillary permeability and ISF composition, creating a compartmental mismatch. To diagnose:
- Monitor Biomarkers: Use a multiplexed sensor (if available) or paired blood samples to track local inflammatory markers (e.g., IL-6, CRP) near the wear site.
- Comparative Site Testing: Place identical sensors at two different anatomical sites. Significant, sustained discrepancies in calculated PK parameters suggest a local site effect.
- Histology: In animal studies, perform histopathological analysis of the tissue under the sensor post-study to grade inflammatory response.
Experimental Protocols for Key Cited Studies
Protocol 1: In Vitro Assessment of Biofouling-Induced Signal Decay Objective: Quantify the rate of signal attenuation due to protein adsorption on a biosensor membrane. Materials: Target biosensor, flow cell system, artificial interstitial fluid (aISF), 4-10 g/L Bovine Serum Albumin (BSA) in aISF (fouling solution), analyte standards. Methodology:
- Calibrate the sensor in fresh aISF with a standard curve (e.g., 5-point calibration).
- Perfuse the fouling solution over the sensor surface at 1 µL/min for 18 hours at 37°C.
- Every 3 hours, stop the fouling solution and perfuse a mid-range analyte standard in clean aISF. Record the sensor response.
- Calculate the percentage response recovery relative to the initial calibration at each time point.
- Plot recovery % vs. time to establish a decay curve for data integrity adjustment.
Protocol 2: In Vivo Validation of Sensor Lag Time in a Rodent PK Study Objective: Determine the time lag between plasma and ISF concentration measurements from a wearable sensor. Materials: Animal model, implantable biosensor, venous catheter, analytical instrument (e.g., LC-MS/MS). Methodology:
- Administer the drug compound via IV bolus.
- Collect serial blood plasma samples via catheter at pre-defined intervals (e.g., 1, 5, 15, 30, 60, 120... min).
- Record the continuous sensor signal from the ISF concurrently.
- Analyze plasma samples via gold-standard method to establish the true plasma PK curve.
- Use deconvolution or cross-correlation analysis to mathematically determine the time shift required to align the sensor-derived profile with the plasma profile. This shift is the aggregate lag time.
Data Presentation
Table 1: Impact of Biofouling on Key PK Parameters (Simulated In-Vitro Data)
| PK Parameter | Unfouled Sensor Value | After 24h Fouling | % Deviation | Clinical Impact |
|---|---|---|---|---|
| C_max (ng/mL) | 125.0 | 98.5 | -21.2% | Underdosing potential |
| T_max (h) | 2.0 | 2.8 | +40.0% | Mis-timed efficacy |
| AUC_0-∞ (h*ng/mL) | 845.3 | 701.6 | -17.0% | Underestimation of exposure |
| Half-life (h) | 6.5 | 7.9 | +21.5% | Altered clearance perception |
Table 2: Efficacy of Anti-Fouling Coatings in Wearable Sensors
| Coating Material | Mechanism | Signal Retention @ 24h | Reduction in Inflammatory Markers (vs. Uncoated) |
|---|---|---|---|
| Polyethylene Glycol (PEG) | Hydrophilic Steric Barrier | 78% | 30% |
| Zwitterionic Polymer | Electrostatic Hydration Layer | 92% | 65% |
| Hydrogel (Alginate) | Physical Separation & Hydration | 85% | 50% |
| Uncoated Reference | N/A | 54% | 0% (Baseline) |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in PK/PD Sensor Research |
|---|---|
| Artificial Interstitial Fluid (aISF) | Simulates the ionic and chemical environment of skin ISF for in-vitro calibration and fouling studies. |
| Bovine Serum Albumin (BSF) / Fibrinogen | Model proteins for simulating the early-stage biofouling (Vroman effect) on sensor surfaces. |
| Zwitterionic Sulfobetaine Monomer | Key reagent for synthesizing anti-fouling polymer brushes on sensor electrodes via surface-initiated polymerization. |
| Fluorescently-tagged Analytic Analog | Allows for concurrent electrochemical sensing and confocal microscopy visualization of analytic diffusion through fouling layers. |
| Microdialysis System | Gold-standard reference method for continuous, minimally diluted ISF sampling to validate sensor accuracy in vivo. |
| Kinase/Phosphatase Activity Reporters | For PD studies: Encapsulated reporters in hydrogel coatings can provide localized, continuous data on drug target engagement. |
Visualizations
Diagram 1: Biofouling Impact on PK/PD Data Pathway
Diagram 2: Sensor Data Integrity Validation Workflow
Technical Support & Troubleshooting Center
FAQ & Troubleshooting Guide
Q1: Our electrochemical sensor shows significant signal drift during prolonged (>8 hour) sweat monitoring. What are the primary culprits and mitigation strategies?
A: Signal drift in sweat sensors is commonly caused by biofouling and changing electrolyte composition. Key culprits are:
- Protein Adsorption: Accumulation of albumin, lysozyme, and immunoglobulins from sweat/ISF on the electrode surface, blocking active sites.
- Microbial Adhesion: Early-stage biofilm formation by skin commensals (e.g., Staphylococcus epidermidis, Cutibacterium acnes).
- Salt Precipitation: Crystallization of NaCl, KCl, and lactate upon sweat evaporation, altering local conductivity.
Mitigation Protocols:
- Surface Pre-conditioning: Soak sensor in artificial sweat (see Table 1) for 1 hour prior to calibration to pre-adsorb proteins.
- Apply Anti-fouling Membranes: Spin-coat a ~100 nm layer of zwitterionic polymer (e.g., poly(sulfobetaine methacrylate)) on the electrode. Protocol: 2% w/v solution in methanol, spin at 3000 rpm for 60 seconds, cure at 60°C for 2 hours.
- Dynamic Baseline Correction: Implement an electrochemical impedance spectroscopy (EIS) measurement at 100 Hz every 15 minutes to track fouling, using the impedance change to correct the amperometric or potentiometric signal.
Q2: How do we differentiate between signal artifacts caused by the skin microbiome versus those from inherent interstitial fluid (ISF) dynamics?
A: This requires controlled in-vitro and on-skin experiments.
Experimental Protocol:
- In-vitro Bioreactor Test:
- Setup: Use a flow cell with your sensor. Prepare three test solutions: (A) Sterile artificial ISF, (B) Artificial ISF with 1 mg/mL bovine serum albumin (BSA), (C) Artificial ISF with BSA and S. epidermidis at 10^5 CFU/mL.
- Procedure: Flow each solution over the sensor at 0.1 µL/min (simulating perspiration) for 24 hours. Record sensor output and perform EIS every 2 hours.
- Analysis: Compare the rate of signal decay and impedance increase. A marked shift in Condition C indicates microbiome-specific fouling, often characterized by a two-stage impedance rise (protein adhesion followed by bacterial adhesion).
- On-Skin Control Experiment:
- Site Preparation: Test on two adjacent forearm sites. Clean both with 70% ethanol. Apply a broad-spectrum antiseptic (e.g., chlorhexidine 2%) to one site and cover for 10 minutes, then rinse with sterile water. The other site receives only ethanol cleaning.
- Sensor Deployment: Apply identical sensors to both sites for 12 hours.
- Analysis: A statistically significant difference in signal stability or drift between the two sites points to a substantial microbiome-derived artifact.
Q3: What is the recommended protocol for collecting and characterizing "authentic" sweat and ISF for in-vitro sensor calibration?
A: Sweat Collection: Use a validated macro-patch method (e.g., PharmChek sweat patch) over 30 minutes during moderate exercise (cycling at 60% VO₂ max). Elute analytes from the patch using 2 mL of 10 mM PBS with 0.1% BSA. Filter through a 0.22 µm PES filter. ISF Collection: Use minimally invasive microneedle (hollow or hydrogel-forming) arrays applied for 30-40 minutes. Centrifuge microneedles to recover ISF (typically 5-20 µL). Characterization: Immediately analyze pH (target range: 4.5-7.0), osmolarity (target: 20-180 mOsm/kg for sweat, ~290 mOsm/kg for ISF), and key ionic strength (Na⁺, K⁺, Cl⁻) via ion chromatography. Aliquot and store at -80°C.
Q4: Our wearable's adhesive fails (detaches or causes irritation) during multi-day studies involving profuse sweating. What material solutions exist?
A: Adhesive failure is a critical interface issue.
- For Detachment: Switch to a moisture-absorbing, breathable adhesive like a polyurethane-based hydrogel adhesive (e.g., Tegaderm HP). It manages sweat by absorbing water and transmitting water vapor. Ensure a moisture vapor transmission rate (MVTR) > 800 g/m²/24hr.
- For Skin Irritation: Implement a layered approach:
- Skin Interface: Use a soft, silicone-based adhesive (e.g., Dow Corning 7-9800). It is gentle and creates a protective barrier.
- Sweat Management: Above it, place a thin, porous hydrocolloid layer to absorb sweat and ISF transudate.
- Device Attachment: Use a stronger acrylic adhesive to attach the sensor housing to the top hydrocolloid layer.
- Always conduct a 48-hour skin patch test on a small cohort before full study deployment.
Table 1: Composition of Common Artificial Biofluids for Sensor Testing
| Component | Artificial Eccrine Sweat (pH 5.5) | Artificial Interstitial Fluid (pH 7.4) | Primary Function in Testing |
|---|---|---|---|
| NaCl | 30 mM | 110 mM | Primary ionic conductor, major osmolyte |
| KCl | 10 mM | 4 mM | Key electrolyte, impacts Nernst potential |
| Lactic Acid | 25 mM | 1-3 mM | Key metabolite, can chelate metals |
| Urea | 5 mM | 4-8 mM | Metabolite, can form hydrogen bonds |
| Glucose | 0.1 mM | 5 mM | Primary analyte for many biosensors |
| BSA (Bovine Serum Albumin) | 0.5 mg/mL | 40 mg/mL | Simulates protein fouling |
| NH₄Cl | 5 mM | - | Simulates ammonium in sweat |
| NaHCO₃ | - | 25 mM | pH buffer (ISF) |
| NaH₂PO₄/Na₂HPO₄ | 1 mM | 1 mM | pH buffer |
Table 2: Common Interferents & Their Typical Concentrations at Skin Interface
| Interferent | Sweat Concentration Range | ISF Concentration Range | Primary Impact on Sensor |
|---|---|---|---|
| Ascorbic Acid | 10-150 µM | 30-100 µM | Oxidizable, causes anodic current artifact |
| Uric Acid | 10-70 µM | 150-500 µM | Oxidizable, causes anodic current artifact |
| Cortisol | 1-50 nM | 50-500 nM | Can adsorb non-specifically |
| Ethanol | Variable (0-1 mM) | Correlated to blood | Affects membrane permeability |
| SDS (from soaps) | Trace to 0.01% | - | Can disrupt lipid membranes on sensors |
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Zwitterionic Monomers (e.g., SBMA) | Create ultra-low fouling hydrogel coatings that resist non-specific protein adsorption via electrostatically induced hydration layers. |
| Poly(ethylene glycol) Dimethacrylate (PEGDMA) | Forms hydrophilic cross-linked networks to create diffusion-limiting or anti-fouling barrier membranes on sensor surfaces. |
| Artificial Sweat/ISF Kits (e.g., Pickering Labs) | Provide standardized, reproducible biofluid simulants for controlled bench-top sensor validation and fouling studies. |
| Live/Dead BacLight Bacterial Viability Kit | Fluorescently stain adhered bacteria on explanted sensor surfaces to quantify and visualize biofilm viability. |
| Protease Inhibitor Cocktail (e.g., EDTA-free) | Added to collected biofluid samples to prevent proteolytic degradation of biomarkers (e.g., peptides, cytokines) during storage. |
| Hydrogel-Forming Microneedle Arrays (e.g., from 3M) | For minimally invasive sampling of ISF to obtain "ground truth" data for calibration of wearable sensor readings. |
| Quartz Crystal Microbalance with Dissipation (QCM-D) | Label-free, real-time measurement of mass (proteins, cells) adsorbing onto sensor material surfaces in liquid. |
Experimental Workflow & Pathway Diagrams
Engineering Solutions: Advanced Methodologies for Antifouling and Stable Sensor Design
Technical Support Center: Troubleshooting & FAQs
Frequently Asked Questions
Q1: My PEGylated sensor surface shows unexpectedly high non-specific protein adsorption after two weeks of storage. What could be the cause? A: This is a common issue related to PEG oxidation and hydrolytic degradation. Poly(ethylene glycol) chains are susceptible to auto-oxidation in the presence of trace metals or UV light, leading to the formation of aldehydes and acids that promote fouling. For storage, ensure the sensor is in an inert atmosphere (e.g., argon-packed container), at -20°C, and with desiccant. Consider adding antioxidants like butylated hydroxytoluene (BHT) to your storage buffer at 0.01-0.1% w/v.
Q2: During zwitterionic polymer brush grafting via ATRP, my polymerization solution becomes viscous and cloudy, resulting in an uneven coating. How can I fix this? A: Cloudiness indicates uncontrolled homogeneous polymerization ("free" polymer in solution) rather than surface-initiated growth. This is typically due to oxygen contamination deactivating the catalyst or an incorrect monomer-to-initiator ratio. Ensure rigorous deoxygenation of all solutions by bubbling with nitrogen or argon for at least 30 minutes prior to use. Confirm your initiator density on the surface; a density that is too low can promote solution-phase polymerization.
Q3: My hydrogel coating significantly attenuates the electrochemical signal of my underlying wearable biosensor. How can I improve signal transduction? A: Signal attenuation is often due to diffusion limitations or excessive hydrogel thickness. You can engineer the hydrogel for better performance by:
- Creating a porosity gradient with a denser, fouling-resistant top layer and a more porous, conductive sub-layer.
- Incorporating conductive nanomaterials (e.g., gold nanoparticles, carbon nanotubes) at ≤ 0.5% w/w into the hydrogel matrix to facilitate electron transfer.
- Reducing the hydrated coating thickness to < 5 µm via spin-coating or electrodeposition techniques.
Q4: How do I quantitatively compare the anti-fouling performance of different surface coatings? A: Standardized quantitative metrics are essential. Use the following table to compare key performance indicators (KPIs):
Table 1: Quantitative Metrics for Anti-Fouling Coating Performance
| Metric | Measurement Technique | Target Value (Excellent Performance) | Typical Range for Effective Coatings |
|---|---|---|---|
| Protein Adsorption | Quartz Crystal Microbalance (QCM-D), Surface Plasmon Resonance (SPR) | < 5 ng/cm² | 5 - 50 ng/cm² |
| Cell Adhesion | Fluorescence microscopy (stained nuclei), Counting | < 100 cells/mm² after 24h | 100 - 500 cells/mm² |
| Δf/ΔD Ratio (QCM-D) | Third overtone frequency (Δf) & dissipation (ΔD) shift | Δf/ΔD < -0.1 Hz/10⁻⁶ | Indicates a rigid, protein-resistant layer |
| Hydration Layer Thickness | Ellipsometry, Neutron Reflectometry | > 10 Å | 10 - 50 Å |
Experimental Protocols
Protocol 1: "Grafting-To" PEGylation on a Gold Sensor Surface Objective: Create a dense monolayer of thiol-terminated PEG to minimize non-specific adsorption.
- Substrate Cleaning: Sonicate gold-coated sensor in ethanol and then in deionized water for 10 minutes each. Dry under N₂ stream.
- Plasma Activation: Treat sensor with oxygen plasma for 2 minutes at 100 W.
- PEG Solution Preparation: Dissolve mPEG-Thiol (MW 5000 Da) in degassed, anhydrous ethanol to a final concentration of 1.0 mM. Add 0.1% v/v triethylamine as a catalyst.
- Immersion Coating: Immerse the clean sensor in the PEG solution for 24 hours at room temperature in a sealed, dark vessel under N₂ atmosphere.
- Rinsing & Storage: Rinse thoroughly with fresh, degassed ethanol and DI water. Dry under N₂. Store under argon at -20°C if not used immediately.
Protocol 2: SI-ATRP of Zwitterionic Poly(sulfobetaine methacrylate) (pSBMA) Objective: Grow a dense, hydrophilic polymer brush coating via surface-initiated atom transfer radical polymerization.
- Surface Initiator Immobilization: Functionalize substrate (e.g., SiO₂, Au) with an ATRP initiator (e.g., (3-aminopropyl)triethoxysilane followed by reaction with 2-bromoisobutyryl bromide).
- Monomer Solution: Dissolve SBMA monomer (2.0 M) and triethylamine (2.2 M) in a 3:1 v/v mixture of methanol and deionized water.
- Deoxygenation: Transfer solution to a Schlenk flask. Freeze with liquid N₂, evacuate, and thaw under N₂. Repeat 3 cycles.
- Catalyst Addition: In a glovebox, add Cu(I)Br and ligand (e.g., HMTETA) to achieve [Monomer]:[Cu(I)]:[Ligand] = 100:1:1 molar ratio.
- Polymerization: Inject the catalyst-containing solution onto the initiator-functionalized substrate. React for 1-4 hours at room temperature.
- Termination: Remove substrate, rinse copiously with DI water and EDTA solution (50 mM) to remove copper catalyst.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Surface Engineering Experiments
| Item | Function | Example Vendor/Product Code |
|---|---|---|
| mPEG-Thiol (MW 2000-5000 Da) | Forms a biocompatible, protein-resistant monolayer on gold surfaces via Au-S bonds. | Sigma-Aldrich, 729108 |
| Carboxybetaine Acrylamide (CBAA) | Zwitterionic monomer for creating ultra-low fouling hydrogel coatings via radical polymerization. | TCI Chemicals, C3059 |
| (3-Aminopropyl)triethoxysilane (APTES) | Coupling agent to introduce amine groups and initiator sites onto oxide surfaces (SiO₂, TiO₂). | Gelest, SIA0610.0 |
| 2-Bromoisobutyryl Bromide (BiBB) | ATRP initiator precursor for functionalizing amine-coated surfaces. | Sigma-Aldrich, 248921 |
| Cu(I)Br & HMTETA Ligand | Catalyst system for ATRP enabling controlled radical polymerization from surfaces. | Sigma-Aldrich, 468711 & 517259 |
| Poly(ethylene glycol) diacrylate (PEGDA, MW 700) | Crosslinker for forming hydrogels with tunable mesh size via UV photopolymerization. | Sigma-Aldrich, 455008 |
| LAP Photoinitiator | Water-soluble, cytocompatible photoinitiator for UV (365 nm) crosslinking of hydrogels. | Toronto Research Chemicals, L006000 |
Visualizations
Title: Surface Engineering Pathways for Sensor Stability
Title: PEGylation "Grafting-To" Protocol Workflow
Title: SI-ATRP Mechanism for Zwitterionic Brushes
Technical Support & Troubleshooting Center
Frequently Asked Questions (FAQs)
Q1: My graphene-based biosensor shows inconsistent electrochemical signal upon repeated sweat exposure. What could be the cause and how can I fix it?
A: Inconsistent signals often stem from nonspecific protein adsorption (biofouling) or instability of the biorecognition element (e.g., enzyme). Perform the following diagnostic:
- Check Electrode Integrity: Use cyclic voltammetry (CV) in a standard ferricyanide solution. A >20% decrease in peak current suggests physical degradation or fouling layer.
- Test for Fouling: Immerse the sensor in a 10 mg/mL BSA solution for 1 hour, then rerun CV. A significant change in charge transfer resistance (Rct) measured via electrochemical impedance spectroscopy (EIS) confirms biofouling.
- Solution: Apply an antifouling nanocoating. A recommended protocol is spin-coating a 2 nmol/mL solution of zwitterionic polymer (e.g., poly(sulfobetaine methacrylate)) grafted with polyethylene glycol (PEG) chains. This creates a hydration layer that reduces protein adhesion by >85%.
Q2: The conductivity of my MXene (Ti₃C₂T₅) film degrades rapidly during prolonged operation in a physiological buffer. How can I improve its stability?
A: MXene oxidation is a common issue. Degradation is characterized by a rise in sheet resistance (>50% over 24 hours) and a color change from metallic to whitish.
- Primary Cause: Reaction with dissolved oxygen and water, leading to the formation of TiO₂.
- Prevention Strategy:
- Encapsulation: Use atomic layer deposition (ALD) to apply a 5-10 nm conformal layer of Al₂O₃. This can extend stable operation lifetime from <24h to >7 days.
- Storage: Always store MXene dispersions under argon at -20°C and use within 72 hours of synthesis.
- In-Situ Stabilization: Incorporate 1 mM ascorbic acid into your measurement buffer as an antioxidant.
Q3: The antifouling polymer brush coating I applied is preventing the adhesion of my capture antibodies. How do I achieve both antifouling and biofunctionalization?
A: This is a challenge of balancing repellency and specific binding. You need a co-modification strategy.
- Solution: Use a mixed self-assembled monolayer (SAM) or a block copolymer approach.
- Protocol: For a gold electrode, incubate in a mixed ethanolic thiol solution containing 90% carboxy-terminated thiol (e.g., 11-mercaptoundecanoic acid) and 10% oligo(ethylene glycol)-terminated thiol (e.g., EG6-thiol) for 24 hours. Rinse. Then, use EDC/NHS chemistry to covalently immobilize antibodies only onto the carboxy-terminated sites. The EG6-thiol domains provide antifouling properties, reducing nonspecific adsorption by ~70-90% while maintaining antibody activity.
Q4: My nanostructured sensor shows excellent sensitivity in buffer but poor performance in complex biofluids (e.g., undiluted serum). What are the next steps?
A: This directly relates to your thesis on sensor stability. The issue is matrix interference.
- Systematic Troubleshooting Guide:
- Validate Assay: Run a calibration curve in the target biofluid. A maintained linear range but shifted baseline indicates fouling. A lost linear range suggests interference with the signaling mechanism.
- Implement a Barrier: Introduce a size-exclusion nanomembrane (e.g., a porous alumina layer with 5-10 nm pores) or a dense hydrogel mesh (e.g., 5% polyacrylamide) above the sensing layer. This physically blocks large proteins and cells while allowing analyte diffusion.
- Data Correction: Use a built-in internal reference electrode (e.g., a bare Au electrode with a known redox couple) to differentiate between signal drift and specific analyte signal.
Key Experimental Protocols
Protocol 1: Synthesis and Antifouling Functionalization of Graphene Oxide (GO) for Sensor Interfaces
Objective: To create a stable, antifouling GO-based platform for biosensing. Materials: Graphite powder, NaNO₃, concentrated H₂SO₄, KMnO₄, H₂O₂ (30%), HCl, poly-L-lysine-grafted-polyethylene glycol (PLL-g-PEG), phosphate-buffered saline (PBS). Steps:
- GO Synthesis (Modified Hummers' Method): In an ice bath, mix 1 g graphite + 0.5 g NaNO₃ in 23 mL concentrated H₂SO₄. Slowly add 3 g KMnO₄ while keeping temp <20°C. Stir at 35°C for 2h. Slowly add 46 mL deionized (DI) water, then heat to 98°C for 15 min. Dilute with 140 mL DI water, add 2.5 mL H₂O₂ (turns mixture yellow). Wash with 1M HCl and DI water via centrifugation until pH ~6.
- Sensor Deposition: Drop-cast 10 µL of 2 mg/mL GO dispersion onto cleaned ITO/Au electrode. Dry at 60°C.
- Antifouling Coating: Incubate GO-coated electrode in 1 mg/mL PLL-g-PEG solution in PBS for 1 hour. Rinse with PBS. The cationic PLL backbone adsorbs to negatively charged GO, presenting a dense PEG brush layer.
Protocol 2: Fabrication of an MXene (Ti₃C₂T₅)-Polymer Composite for Flexible Electrodes
Objective: To produce a flexible, oxidation-resistant MXene electrode for wearable applications. Materials: Ti₃AlC₂ MAX phase, LiF, HCl, DI water, polyurethane (PU) dispersion, vacuum filtration setup. Steps:
- MXene Etching: Slowly add 1 g LiF to 20 mL of 9M HCl while stirring. Add 1 g Ti₃AlC₂ powder over 10 min. Etch at 35°C for 24h under stirring. Wash with DI water via centrifugation (3500 rpm, 5 min cycles) until supernatant pH >6. Decant and collect the multilayered sediment.
- Delamination: Resuspend sediment in DI water and probe-sonicate under Ar for 1h. Centrifuge at 3500 rpm for 1h; collect the dark colloidal supernatant (single/few-layer MXene).
- Composite Formation: Mix MXene dispersion with PU dispersion at a 3:1 weight ratio (MXene:PU). Stir for 12h.
- Film Casting: Vacuum filter the composite mixture onto a porous PVDF membrane. Air-dry, then peel off to obtain a free-standing, flexible film (~5-10 µm thick).
Table 1: Comparison of Nanomaterial Antifouling Performance in Human Serum
| Nanomaterial Platform | Coating/Modification | % Reduction in Nonspecific Protein Adsorption (vs. Bare Au) | Measurement Technique | Reference Stability (Hours) |
|---|---|---|---|---|
| Graphene (CVD) | Zwitterionic Peptide | 92% | Quartz Crystal Microbalance (QCM) | 72 |
| Reduced GO | PEGylated Lipid Bilayer | 88% | Surface Plasmon Resonance (SPR) | 48 |
| MXene (Ti₃C₂T₅) | Native (No Coating) | 45% | Fluorescence Microscopy | <12 |
| MXene (Ti₃C₂T₅) | ALD Al₂O₃ (10 nm) + Heparin Gel | 91% | Electrochemical Impedance Spectroscopy (EIS) | 168 |
| Gold Nanorods | Poly(sulfobetaine) Brush | 95% | QCM-D | 96 |
Table 2: Key Sensor Performance Metrics with Antifouling Nanostructures
| Sensor Type | Target Analyte | Limit of Detection (in Buffer) | Limit of Detection (in Serum) | Signal Drift over 10h (in Serum) | Key Nanomaterial & Strategy |
|---|---|---|---|---|---|
| Electrochemical | Glucose | 2.1 µM | 5.8 µM | 4.2% | Graphene/PEG-yne Network |
| Electrochemical | Cortisol | 0.8 nM | 2.5 nM | 8.7% | MXene/Molecularly Imprinted Polymer |
| Optical (SPR) | CRP | 15 ng/mL | 42 ng/mL | 6.1% | Au Nanodisk/Zwitterionic Hydrogel |
| Field-Effect Transistor (FET) | Dopamine | 100 pM | 1.2 nM | 12.5% | Graphene/Phospholipid Bilayer |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function/Description |
|---|---|
| PLL(20)-g[3.5]-PEG(5) | Poly-L-lysine grafted with polyethylene glycol. Cationic backbone adsorbs to negative surfaces, presenting a dense, protein-repellent PEG brush layer. |
| HS-(CH₂)₁₁-EG₆-OH | Thiol-terminated oligo(ethylene glycol) alkanethiol. Forms self-assembled monolayers on gold, creating a highly ordered, hydrophilic antifouling surface. |
| DSPE-PEG(2000)-Biotin | 1,2-distearoyl-sn-glycero-3-phosphoethanolamine conjugated to PEG and biotin. Used to create functionalizable lipid bilayers on nanostructures; PEG provides antifouling, biotin allows streptavidin-antibody linkage. |
| Carboxylated MXene Quantum Dots | Ultra-small, water-dispersible MXene fragments with -COOH groups. Enhance electron transfer in composites and provide sites for covalent biomolecule immobilization. |
| Zwitterionic Sulfobetaine Silane | A silane coupling agent bearing zwitterionic groups. Forms a durable, covalently attached super-hydrophilic monolayer on oxide surfaces (SiO₂, ITO) to resist cell and protein adhesion. |
Experimental Workflow & Pathway Diagrams
Title: Troubleshooting Biosensor Signal Degradation Workflow
Title: Nanomaterial Antifouling Strategies and Mechanisms
Title: Fabrication of Stable MXene Composite Electrode Protocol
Technical Support Center: Troubleshooting & FAQs
FAQ & Troubleshooting Guide
Q1: My fabricated pH-responsive hydrogel coating does not exhibit reversible swelling/deswelling. What could be the cause? A: This is often due to insufficient crosslinking or incorrect monomer ratio. Ensure your crosslinker (e.g., N,N'-methylenebisacrylamide) concentration is between 0.5-2.0 mol% relative to the primary monomer (e.g., acrylic acid for anionic response). Verify the polymerization initiator (e.g., APS) is fresh and the reaction proceeded under inert atmosphere (N₂) to prevent oxygen inhibition.
Q2: My thermoresponsive polymer brush surface (e.g., pNIPAM) shows inconsistent lower critical solution temperature (LCST) behavior and poor anti-biofouling performance. How can I fix this? A: Inconsistent LCST often stems from polydispersity or residual monomer. Repurify the polymer via dialysis (MWCO 3.5 kDa) against cold DI water for 72 hours, changing water every 12 hours. For brushes, ensure the surface initiator density (via SI-ATRP) is optimized—target 0.2-0.4 chains/nm². Low density reduces switching efficacy.
Q3: The "self-cleaning" property of my lotus-inspired superhydrophobic surface is lost after abrasion or protein exposure. How do I improve durability? A: Superhydrophobicity relies on hierarchical micro/nano-structures which are fragile. Incorporate a durable polymeric binder (e.g., perfluorinated polyurethane) into your nanoparticle (SiO₂, TiO₂) coating solution. Alternatively, design a self-healing surface by embedding microcapsules containing fluoroalkyl silanes (e.g, 1H,1H,2H,2H-perfluorodecyltriethoxysilane) that rupture upon scratch.
Q4: My electroresponsive conducting polymer (e.g., PEDOT:PSS) film delaminates from the wearable sensor substrate during cyclic voltage application. A: Delamination indicates poor adhesion. Pre-treat your flexible substrate (e.g., PDMS, polyimide) with a silane adhesion promoter (3-aminopropyltriethoxysilane) or apply a thin primer layer of PEDOT:PSS mixed with 3-glycidyloxypropyltrimethoxysilane (GOPS) crosslinker at 1:0.03 v/v ratio. Curing at 140°C for 15 minutes enhances bonding.
Q5: The enzymatic biofouling release efficacy of my UV-light-responsive azobenzene surface is below 40% in complex biofluids (e.g., sweat, interstitial fluid). A: Complex fluids contain proteins that form a dense, multilayer foulant. Augment the photoresponse with a synergistic zwitterionic polymer underlayer (e.g., poly(sulfobetaine methacrylate)). The azobenzene provides topological change, while the zwitterion provides a hydration barrier. Use UVA light at 365 nm, 5 mW/cm² for 10 min cycles.
Quantitative Performance Data Table
Table 1: Comparison of Stimuli-Responsive Coating Performances for Biofouling Mitigation in Wearable Biosensors
| Coating Type | Stimulus | Response Time | Biofouling Reduction (% vs Control) | Cycling Stability (# of cycles) | Key Limitation |
|---|---|---|---|---|---|
| pNIPAM Brushes | Temperature (Δ~10°C) | 30-90 seconds | 60-75% (Protein) | 50-100 | Slow in viscous media |
| pH-responsive Hydrogel | pH (5.0 to 7.4) | 2-5 minutes | 55-70% (Cells) | 20-50 | Ionic strength sensitivity |
| Superhydrophobic SiO₂/ Fluoropolymer | Mechanical (Shear) | Instantaneous | 70-85% (Bacteria) | 10-30 before wear | Poor mechanical durability |
| Azobenzene-PEG | UV Light (365 nm) | 1-2 minutes | 40-60% (Complex Biofluid) | 100+ | Potential UV damage to skin/sample |
| Conducting Polymer (PEDOT) | Electric Field (±0.5 V) | 5-30 seconds | 65-80% (Protein) | 200+ | Requires integrated electrodes |
Detailed Experimental Protocols
Protocol 1: Fabrication of Durable Superhydrophobic Coating for Wearable Sensor Patches Objective: Create an abrasion-resistant, lotus-leaf-inspired coating to prevent adhesion of biological fluids and contaminants. Materials: Fumed silica nanoparticles (7-40 nm), 1H,1H,2H,2H-Perfluorooctyltriethoxysilane (PFOTES), Ethanol (absolute), Polyurethane dispersion (anionic, 30% solids), Acetic acid (glacial). Procedure:
- Hydrophobization of Nanoparticles: Disperse 2g fumed silica in 100 mL ethanol. Add 4 mL PFOTES and 2 mL DI water. Stir at 60°C for 6 hours. Centrifuge, wash with ethanol, and dry.
- Coating Formulation: Dissolve 5g polyurethane in 50 mL ethanol/water (4:1) mix. Add 0.5g hydrophobized silica. Sonicate for 30 min. Add 0.1 mL acetic acid to catalyze crosslinking.
- Application: Spray-coat (0.2 MPa, 20 cm distance) or dip-coat onto pre-cleaned sensor substrate (e.g., PET).
- Curing: Cure at 120°C for 1 hour. Characterize water contact angle (>150°) and sliding angle (<10°).
Protocol 2: Grafting pH-Responsive Hydrogel for On-Demand Drug Release in Wearable Patches Objective: Synthesize a poly(methacrylic acid-co-poly(ethylene glycol) dimethacrylate) hydrogel that swells at high pH to release an antimicrobial agent (e.g., chlorhexidine). Materials: Methacrylic acid (MAA), Poly(ethylene glycol) dimethacrylate (PEGDMA, Mn 750), 2-Hydroxy-2-methylpropiophenone (photoinitiator), Phosphate Buffered Saline (PBS, pH 7.4 & 5.0). Procedure:
- Pre-gel Solution: Mix MAA (85 µL), PEGDMA (15 µL), and photoinitiator (2 wt% of total monomers) in 1 mL DI water. Degas with N₂ for 5 min.
- Photopolymerization: Inject solution into a mold (e.g., 100 µm spacer between glass slides). Expose to UV light (320-390 nm, 10 mW/cm²) for 3 minutes.
- Swelling/Release Test: Hydrate gel in PBS pH 5.0. Measure initial mass (Mdry). Transfer to PBS pH 7.4, measure mass over time (Mwet). Swelling Ratio = (Mwet - Mdry)/M_dry. For release, load hydrogel with 1 mg/mL drug in step 1 and measure UV-Vis absorbance of supernatant after pH switch.
Visualizations
Workflow: Developing Self-Cleaning Wearable Sensor Interfaces
pNIPAM Thermal Response & Biofouling Release Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Stimuli-Responsive Surface Experiments
| Reagent/Material | Function/Application | Example Supplier(s) |
|---|---|---|
| N-Isopropylacrylamide (NIPAM) | Monomer for thermoresponsive polymers (LCST ~32°C). Purify by recrystallization. | Sigma-Aldrich, TCI |
| (3-Aminopropyl)triethoxysilane (APTES) | Coupling agent for anchoring polymers/nanoparticles to oxide surfaces (SiO₂, TiO₂). | Gelest, Merck |
| 1H,1H,2H,2H-Perfluorodecyltrichlorosilane (FDTS) | Creates low-surface-energy monolayer for superhydrophobic surfaces. | ABCR, Sigma-Aldrich |
| Poly(3,4-ethylenedioxythiophene):poly(styrene sulfonate) (PEDOT:PSS) | Conductive polymer for electroresponsive coatings and electrodes. | Heraeus, Ossila |
| Azobenzene-4,4'-dicarboxylic acid | Photoswitchable molecule for UV-light-triggered topological changes. | Tokyo Chemical Industry |
| Poly(sulfobetaine methacrylate) (pSBMA) | Zwitterionic polymer for creating a hydration barrier against non-specific adsorption. | Specific polymers, self-synthesized |
| Poly(ethylene glycol) diacrylate (PEGDA, Mn 700) | Crosslinker for hydrophilic, protein-resistant hydrogel networks. | Polysciences, Sigma-Aldrich |
| Silanized Silica Nanoparticles (10-20 nm) | Building blocks for creating durable, hierarchical rough surfaces. | Nanocs, US Research Nanomaterials |
Technical Support Center
Troubleshooting Guide & FAQs
Q1: During in vitro testing, my sensor signal shows an initial sharp peak followed by a rapid, non-reproducible decay. What could be the cause? A: This is a classic symptom of protein biofouling and membrane saturation. The initial peak corresponds to the target analyte reaching the transducer surface. The rapid decay indicates that non-specific adsorption of proteins (e.g., albumin, fibrinogen) is blocking the active sites or impeding diffusion through the outermost membrane. This fouling layer alters local hydrodynamics and permeability.
- Solution:
- Verify Membrane Integrity: Inspect the diffusion-limiting membrane for pinholes or delamination using SEM if possible.
- Implement a Pre-conditioning Step: Soak the sensor in a solution mimicking the sample matrix (e.g., 1% BSA in PBS) for 1-2 hours prior to calibration to passivate non-specific sites.
- Re-evaluate Membrane Hydrophilicity: Consider applying an additional cross-linked hydrogel layer (e.g., PEGDA) to enhance antifouling properties. Refer to Protocol 1.
Q2: My multi-layer sensor exhibits high sensitivity in buffer but fails in complex biofluids (e.g., sweat, ISF). How can I diagnose the layer-by-layer failure? A: This indicates a failure in the hierarchical design's selectivity. The issue likely lies in the interference-rejection layer or the diffusion-limiting membrane's performance in a fouling medium.
- Solution:
- Perform a Selectivity Audit: Test the sensor against common interferents (e.g., ascorbic acid, uric acid, lactate, acetaminophen) individually in buffer. See Table 1 for typical concentration ranges.
- Conduct a Layer-by-Layer Characterization:
- Use Electrochemical Impedance Spectroscopy (EIS) to monitor the charge-transfer resistance (Rct) after exposure to each layer-deposition step and after exposure to biofluid.
- A jump in Rct after biofluid exposure suggests successful blocking by the antifouling layer. No change suggests layer failure.
- Optimize the Diffusion-Limiting Membrane: The linear range may be mismatched. Thicken the membrane to increase the diffusion path or adjust its porosity. Refer to Protocol 2.
Q3: The baseline drift of my wearable sensor exceeds 5% per hour during continuous monitoring. What are the primary mechanical and electrochemical culprits? A: Excessive baseline drift compromises long-term stability and is often multi-factorial.
- Solution:
- Check Mechanical Stability: Ensure the multi-layer architecture is securely laminated. Delamination creates micro-environments that concentrate analytes or buffer pH.
- Electrochemical Leakage: Verify the integrity of the reference electrode. In solid-contact designs, leakage of internal electrolyte can cause drift. Consider using a more stable reference membrane (e.g., PVC-based ion-selective membrane containing a lipophilic salt).
- Membrane Hydration Swelling: If using hydrogel-based layers, pre-hydrate the sensor for a consistent period (e.g., 30 mins in PBS) before baseline measurement to reach swelling equilibrium.
Q4: When integrating a new anti-biofouling polymer (e.g., zwitterionic hydrogel) into my multi-layer stack, adhesion failure occurs. How can I improve interlayer adhesion? A: Adhesion failure between dissimilar materials (e.g., metal electrode, hydrophobic membrane, hydrophilic hydrogel) is common.
- Solution:
- Surface Functionalization: Use oxygen plasma treatment on polymer membranes (30-60 seconds, 50-100W) to create -OH groups for better bonding with hydrophilic layers.
- Use an Adhesion Promoter: Apply a thin layer of a silane coupling agent (e.g., (3-aminopropyl)triethoxysilane for oxide surfaces) or a dopamine hydrochloride solution (2 mg/mL in 10 mM Tris buffer, pH 8.5, 30 min coating) to create a universal primer layer.
- Cross-linking Strategy: Employ a cross-linker that reacts with functional groups on both adjacent layers (e.g., glutaraldehyde for amine-containing layers).
Research Reagent Solutions Toolkit
| Item | Function & Rationale |
|---|---|
| Poly(ethylene glycol) diacrylate (PEGDA, MW 700) | A photopolymerizable hydrogel precursor. Forms a highly hydrophilic, cross-linked network that reduces non-specific protein adsorption by creating a hydration barrier. |
| 2-Hydroxy-2-methylpropiophenone (Photoinitiator) | Used with PEGDA. Generates free radicals under UV light (365 nm) to initiate cross-linking polymerization. |
| Polyurethane (e.g., ChronoFlex AR) | A common diffusion-limiting membrane material. Provides tunable permeability and mechanical robustness for controlling analyte flux to the transducer. |
| Nafion Perfluorinated Resin | A cation-exchange polymer coating. Used as an interference-rejection layer to repel anionic interferents (e.g., ascorbate, urate) based on charge exclusion. |
| Poly(3,4-ethylenedioxythiophene) Polystyrene sulfonate (PEDOT:PSS) | A conductive polymer. Used as a solid-contact layer in potentiometric sensors to enhance charge capacity and stabilize the potential at the transducer interface. |
| Phosphate Buffered Saline (PBS) with 1% Bovine Serum Albumin (BSA) | Standard pre-conditioning and stability-testing solution. Mimics the proteinaceous matrix of biofluids for in vitro fouling experiments. |
| Dopamine Hydrochloride | Used to form a polydopamine adhesion primer layer on virtually any substrate, promoting binding for subsequent layers. |
| (3-Aminopropyl)triethoxysilane (APTES) | A silane coupling agent. Creates amine-terminated surfaces on silicon/glass or metal oxides for covalent attachment of next layer. |
Table 1: Common Interferents in Biofluids & Sensor Rejection Targets
| Interferent | Typical Concentration in Sweat | Typical Concentration in ISF | Target Rejection Ratio (Signal Change) |
|---|---|---|---|
| Ascorbic Acid | 10 - 150 µM | 20 - 100 µM | < ±2% for 100 µM interferent |
| Uric Acid | 20 - 700 µM | 100 - 500 µM | < ±2% for 500 µM interferent |
| Lactate | 5 - 60 mM | 1 - 15 mM | Must not cross-react with glucose oxidase |
| Acetaminophen | N/A (systemic) | 10 - 200 µM (therapeutic) | < ±5% for 200 µM interferent |
Table 2: Impact of Membrane Layers on Key Sensor Metrics
| Architecture Layer | Primary Function | Target Impact on Sensitivity | Target Impact on Response Time (t90) | Effect on Fouling (ΔSignal after 24h) |
|---|---|---|---|---|
| Base Transducer | Signal generation | Reference (100%) | < 5 s | > -50% (Severe fouling) |
| + Conductive Polymer (PEDOT) | Stability & Charge Capacity | ±5% | + 1-3 s | > -40% |
| + Interference Layer (Nafion) | Selectivity | -10 to -20% | + 5-10 s | > -30% |
| + Diffusion-Limiting Membrane (PU) | Linear Range Control | -30 to -50% | + 15-60 s | > -20% |
| + Anti-fouling Hydrogel (PEGDA) | Biofouling Resistance | ±5% (of final signal) | + 5-15 s | < -5% (Target) |
Experimental Protocols
Protocol 1: Fabrication of a PEGDA-based Anti-biofouling Hydrogel Layer via UV Cross-linking
Objective: To apply a hydrophilic, protein-resistant topcoat on a sensor surface.
- Surface Preparation: Clean the sensor substrate (e.g., gold electrode, polymer membrane) with ethanol and deionized water. Dry under N₂ stream.
- Solution Preparation: Prepare a pre-polymer solution containing 20% (v/v) PEGDA (700 MW) and 1% (v/v) photoinitiator (2-hydroxy-2-methylpropiophenone) in DI water. Vortex for 1 minute.
- Deposition: Pipette 10 µL of the solution onto the active sensor area.
- Cross-linking: Expose the coated sensor to UV light (365 nm, 10 mW/cm²) for 60 seconds in an inert (N₂) atmosphere to prevent inhibition by oxygen.
- Post-processing: Rinse the sensor gently with DI water for 30 seconds to remove unreacted monomers. Soak in PBS (pH 7.4) for 1 hour to hydrate the hydrogel before testing.
Protocol 2: Optimizing a Polyurethane Diffusion-Limiting Membrane by Spin-Coating
Objective: To deposit a reproducible, thin polymer membrane for controlled analyte diffusion.
- Polymer Solution Prep: Dissolve medical-grade polyurethane (e.g., ChronoFlex AR) at 3% (w/v) in a 70:30 mixture of tetrahydrofuran (THF) and dimethylformamide (DMF). Stir overnight.
- Substrate Mounting: Secure the sensor chip (with underlying layers complete) onto a spin coater vacuum chuck.
- Deposition Cycle:
- Dispense: Pipette 50 µL of polymer solution onto the center of the static chip.
- Spread: Spin at 500 RPM for 5 seconds (acceleration 1000 rpm/s).
- Thin: Immediately spin at a final speed (e.g., 2000-5000 RPM, optimized for desired thickness) for 30 seconds.
- Curing: Transfer the coated sensor to a vacuum desiccator for 4 hours to remove residual solvents.
- Characterization: Measure membrane thickness using a profilometer. Calibrate sensor performance in standard analyte solutions to correlate thickness/porosity with linear range and response time.
Diagrams
Diagram Title: Multi-Layer Sensor Architecture for Biofouling Resistance
Diagram Title: Layer-by-Layer Sensor Failure Diagnosis Workflow
Integration with Microfluidics for Continuous Sample Renewal and Calibration
Technical Support Center: Troubleshooting & FAQs
Q1: During continuous operation, my microfluidic flow becomes irregular and eventually stops. What could be the cause? A: This is a classic symptom of biofouling or particle clogging within the microchannels, especially near the sensor interface. First, inspect the waste reservoir for backpressure. Implement the following protocol: 1) Flush with 0.5% (w/v) sodium dodecyl sulfate (SDS) solution for 30 minutes at 20 µL/min. 2) Rinse with deionized water for 15 minutes. 3) Perform a calibration run with fresh buffer. If the problem persists, the issue may be with the peristaltic pump tubing; check for wear and replace if necessary.
Q2: My calibration injections are not producing stable sensor plateaus. The signal drifts during the calibration phase. A: This indicates a failure in achieving a steady-state concentration at the sensor surface, often due to improper flow rate or mixing. Ensure your flow rate is optimized for your channel geometry to allow for complete diffusion. As a rule of thumb, the flow rate (Q, in µL/min) should be less than (D * A) / (L * 10), where D is the analyte diffusion coefficient (~10⁻⁶ cm²/s for glucose), A is cross-sectional area (mm²), and L is channel length (mm) to the sensor. Verify using the following table of recommended starting flow rates:
Table 1: Recommended Flow Rates for Calibration Stability
| Channel Height (µm) | Channel Width (µm) | Target Analyte | Recommended Flow Rate (µL/min) for Steady-State |
|---|---|---|---|
| 100 | 200 | Glucose | 0.5 - 2.0 |
| 100 | 200 | Lactate | 0.5 - 2.0 |
| 50 | 100 | Cortisol | 0.1 - 0.5 |
| 150 | 300 | Sodium Ions | 1.0 - 3.0 |
Q3: How can I verify that my "continuous sample renewal" is effectively reducing biofouling in my wearable sensor experiment? A: Implement a comparative protocol with a fluorescent dye or labeled protein (e.g., FITC-BSA at 0.1 mg/mL) as a fouling marker. Protocol:
- Setup: Operate the microfluidic system with a physiologically relevant buffer (e.g., artificial interstitial fluid) spiked with 0.1 mg/mL FITC-BSA.
- Static Control: Stop flow for 60 minutes over the sensor region. Measure fluorescence intensity (I_static) at the sensor surface via confocal microscopy or integrated photodetector.
- Continuous Flow Test: Re-establish flow at 3 µL/min for 60 minutes. Measure fluorescence intensity (I_flow).
- Calculation: Calculate the Fouling Reduction Ratio (FRR) = (Istatic - Iflow) / I_static * 100%. An FRR > 70% indicates effective renewal. Document results as in Table 2.
Table 2: Example Biofouling Reduction Data
| Experiment Condition | Flow Rate (µL/min) | Duration (min) | Avg. Fluorescence Intensity (a.u.) | Calculated FRR (%) |
|---|---|---|---|---|
| Static (Control) | 0 | 60 | 1250 ± 150 | 0 |
| Continuous Renewal | 1.0 | 60 | 650 ± 80 | 48 |
| Continuous Renewal | 3.0 | 60 | 300 ± 40 | 76 |
| Continuous Renewal | 5.0 | 60 | 280 ± 35 | 78 |
Q4: The automated calibration cycle is causing bubbles that interfere with sensor readings. How do I mitigate this? A: Bubbles often form due to temperature changes, peristaltic pump pulsation, or degassing of fluids. Solutions: 1) Place all buffers and samples in a degassing chamber for 15 minutes before the experiment. 2) Install an in-line bubble trap before the microfluidic chip inlet. 3) Add 0.01% (v/v) Tween 20 to your calibration buffers (ensure it does not interfere with sensing chemistry). 4) Program a "soft start" for your pump: ramp to target flow rate over 5 seconds instead of an immediate start.
Q5: What is the optimal frequency for in-line calibration in a week-long wearable study to compensate for drift without consuming excessive reagent? A: Based on recent studies of enzymatic sensor drift in microfluidic wearables, a tiered calibration strategy is most efficient. Initial frequent calibration characterizes the drift profile, followed by less frequent maintenance.
Protocol: Tiered In-Line Calibration Schedule:
- Day 1-2: Perform a full 5-point calibration (0, Low, Mid, High, 0) every 6 hours.
- Day 3-7: Based on initial drift data, reduce to a single-point verification (at Mid concentration) every 12 hours, with a full 5-point calibration every 48 hours.
- Use an algorithm (e.g., linear regression of baseline drift) to adjust data between calibrations.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Microfluidic Integration Experiments
| Item/Reagent | Function & Brief Explanation |
|---|---|
| Polydimethylsiloxane (PDMS) | Elastomeric polymer for rapid prototyping of microfluidic chips via soft lithography; gas-permeable, optically clear. |
| Phosphate Buffered Saline (PBS) with 0.01% Azide | Standard ionic strength buffer for biosensor testing; azide inhibits microbial growth in reservoirs during long-term studies. |
| Artificial Interstitial Fluid (AISF) | Physiologically relevant test matrix containing NaCl, KCl, MgCl₂, CaCl₂, and HEPES buffer at skin ISF pH (~6.8). |
| Fluorinated Ethylene Propylene (FEP) Tubing | Chemically inert, low-protein-adhesion tubing for peristaltic pumps; reduces analyte absorption and fouling vs. standard PVC. |
| FITC-labeled Bovine Serum Albumin (FITC-BSA) | Model fouling protein used to quantify biofouling accumulation on sensor surfaces via fluorescence measurement. |
| Electrochemical Calibration Standards (e.g., for Glucose) | Pre-mixed, certified concentrations of analyte (e.g., 0, 2.5, 5.0, 10.0, 20.0 mM glucose) for generating calibration curves. |
| Anti-biofouling Surface Primer (e.g., PLL-g-PEG) | Poly(L-lysine)-graft-poly(ethylene glycol) solution; forms a hydrophilic, protein-repellent monolayer on sensor surfaces prior to use. |
Experimental Workflow & Signaling Pathway Diagrams
From Lab to Skin: Troubleshooting Field Failures and Optimizing Long-Term Performance
FAQs & Troubleshooting Guides
Q1: After a 72-hour continuous wear study, my glucose sensor shows a persistent positive baseline shift. What is the likely failure mode and how can I confirm it analytically?
A: This is a classic symptom of a Type 1 (Biofouling-Mediated) failure. The shift is likely due to the non-specific adsorption of proteins (e.g., albumin, fibrinogen) and inflammatory cells at the sensor-skin interface, creating a diffusion barrier. To confirm:
- Protocol 1: Surface Fourier Transform Infrared Spectroscopy (FTIR): Clean the sensor membrane with deionized water and air-dry. Acquire spectra in Attenuated Total Reflection (ATR) mode from 4000-600 cm⁻¹. Compare pre- and post-use spectra. Look for new amide I (~1650 cm⁻¹) and amide II (~1550 cm⁻¹) peaks, indicative of protein adsorption.
- Protocol 2: Scanning Electron Microscopy (SEM): Dehydrate the sensor membrane in a graded ethanol series (30%, 50%, 70%, 90%, 100%). Critical point dry, sputter-coat with 5 nm gold-palladium. Image at 5-10 kV. A fouled surface will show a confluent, matte layer or aggregated cellular debris versus the clean, porous membrane structure.
Q2: My lactate sensor's sensitivity dropped by >60% after repeated calibration cycles in artificial sweat. What post-hoc tests can determine if the enzyme layer has degraded?
A: This suggests a Type 2 (Electrochemical Component Degradation) failure, potentially involving lactate oxidase (LOx) denaturation or leaching.
- Protocol 3: Activity Assay via Spectrophotometry: Carefully remove the enzyme membrane and solubilize it in 1 mL of phosphate buffer (0.1 M, pH 7.0). Add 10 µL of the solution to a cuvette containing 1 mL of assay mix (0.1 M phosphate buffer, 0.5 mM homovanillic acid, 4 U/mL horseradish peroxidase, 10 mM lactate). Monitor the increase in absorbance at 315 nm for 2 minutes. Compare the rate to a fresh enzyme membrane control.
- Protocol 4: Electrochemical Impedance Spectroscopy (EIS) of the Working Electrode: Perform EIS on the used sensor in 5 mM K₃[Fe(CN)₆]/K₄[Fe(CN)₆] in PBS. Use a frequency range of 100 kHz to 0.1 Hz at 10 mV amplitude. Fit the Nyquist plot to a modified Randles circuit. A large increase in charge transfer resistance (Rct) indicates a loss of enzymatic activity facilitating electron transfer.
Q3: How can I differentiate between signal drift caused by reference electrode (Ag/AgCl) poisoning versus membrane biofouling?
A: These require distinct analytical pathways. Use the decision protocol below.
Table 1: Key Analytical Techniques for Failure Mode Diagnosis
| Failure Mode Suspected | Primary Technique | Key Quantitative Metric | Expected Shift (Failed vs. Control) | Confirmatory Technique |
|---|---|---|---|---|
| Biofouling (Type 1) | SEM Imaging | Surface Coverage (%) | >40% coverage by foreign material | ATR-FTIR (Amide peak area) |
| Enzyme Degradation (Type 2) | Spectrophotometric Activity Assay | Enzyme Activity (U/mg) | >60% loss of specific activity | EIS (Increase in Rct) |
| Reference Electrode Poisoning | Open Circuit Potential (OCP) | Potential Drift (mV) | Drift > ±15 mV over 5 min | XPS (Atomic % Cl on Ag/AgCl surface) |
| Membrane Delamination | Optical Profilometry | Roughness (Ra, nm) & Delamination Area (µm²) | Ra increase >200%, area >1000 µm² | Cross-sectional SEM |
Experimental Protocol Detail
Protocol 5: X-ray Photoelectron Spectroscopy (XPS) for Reference Electrode Analysis
- Objective: Determine chemical composition changes on the Ag/AgCl electrode surface.
- Method:
- Sample Prep: Gently rinse the used sensor with DI water. Using fine tweezers, carefully detach the reference electrode. Dry under a nitrogen stream.
- Mounting: Affix the electrode to the XPS sample holder using double-sided conductive carbon tape.
- Analysis: Insert into the XPS chamber. Evacuate to <5 x 10⁻⁸ Torr. Use a monochromatic Al Kα X-ray source (1486.6 eV).
- Survey Scan: Acquire over 0-1200 eV binding energy with 1.0 eV step and 100 eV pass energy.
- High-Resolution Scans: Acquire spectra for Ag 3d, Cl 2p, O 1s, C 1s, and N 1s regions with 0.1 eV step and 20 eV pass energy.
- Data Analysis: Use CasaXPS software. Correct charge referencing to C1s at 284.8 eV. Fit peaks using Gaussian-Lorentzian functions. Crucial: Calculate the Cl:Ag atomic ratio. A ratio significantly below 0.3 (theoretical for AgCl is ~1.0) indicates chloride depletion or surface contamination.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Post-Use Characterization
| Item | Function in Analysis | Example Product/Catalog # | Notes |
|---|---|---|---|
| Phosphate Buffered Saline (PBS), 10X, Molecular Biology Grade | Gentle rinsing of biological residues from sensor surface without damaging delicate layers. | ThermoFisher, AM9625 | Always dilute to 1X and adjust to pH 7.4 before use. |
| Homovanillic Acid (HVA) | Fluorogenic substrate used in peroxidase-coupled enzyme activity assays (e.g., for oxidase-based sensors). | Sigma-Aldrich, H1259 | Prepare fresh in DMSO; light sensitive. |
| Horseradish Peroxidase (HRP), Lyophilized | Coupling enzyme for colorimetric/fluorometric detection of H₂O₂ generated by sensor oxidases. | MilliporeSigma, 516531 | Reconstitute in buffer, aliquot, and store at -20°C. |
| Potassium Ferri-/Ferrocyanide | Redox probe for Electrochemical Impedance Spectroscopy (EIS) to assess electrode integrity and fouling. | Sigma-Aldrich, 60279 (K₃Fe(CN)₆) / 455946 (K₄Fe(CN)₆) | Use equimolar 5 mM solution in supporting electrolyte. |
| Critical Point Dryer (CPD) Media | For dehydrating biological samples (e.g., fouled membranes) prior to SEM without structural collapse. | Electron Microscopy Sciences, Liquid CO₂, 16715 | Essential for accurate topographical imaging of hydrated biofilms. |
| Gold/Palladium Sputtering Target | For applying a thin, conductive coating to non-conductive samples (e.g., polymer membranes) for SEM. | Ted Pella, 50/50 Au/Pd, 91111 | 5-10 nm thickness is typically sufficient. |
Technical Support & Troubleshooting Center
Frequently Asked Questions (FAQs)
Q1: During in vitro testing, my coated sensor shows excellent fouling resistance but a severely diminished analyte flux. What is the most likely cause and how can I troubleshoot it? A: The most likely cause is excessive coating thickness or overly dense cross-linking, which increases the diffusion path length and creates a significant mass transfer barrier. To troubleshoot:
- Characterize: Use ellipsometry or profilometry to measure the actual coating thickness. Aim for sub-micron (often 100-500 nm) layers.
- Optimize Deposition: For layer-by-layer (LbL) or spin-coated films, systematically reduce the number of deposition cycles or spin speed/duration.
- Modify Formulation: For hydrogel coatings, slightly increase the water content or porosity by adjusting the monomer-to-crosslinker ratio. Refer to Table 1 for target values.
Q2: My sensor coating performs well in buffer but fails rapidly in complex biofluids (e.g., serum, sweat). What strategies can improve stability? A: This indicates non-specific protein adsorption (fouling) is overwhelming the coating.
- Incorporate Zwitterions: Introduce polymers like poly(sulfobetaine methacrylate) (pSBMA) into your coating matrix. Their strong hydration layer resists protein adhesion.
- Increase Cross-linking Density: Ensure your coating is sufficiently cross-linked to prevent biofilm precursors from penetrating and degrading the matrix.
- Validate with Real Fluids: Always include a final validation step in undiluted, relevant biofluid. See Protocol 1.
Q3: How do I quantitatively measure the permeability (P) and selectivity (β) of my anti-fouling coating? A: Use a diffusion cell apparatus with fluorescently tagged model analytes and interferents.
- Analyte Choice: Use a target molecule (e.g., 1 kDa dextran for glucose mimic) and a similar-sized interferent (e.g., bovine serum albumin, BSA).
- Calculate: Measure the flux (J) across the coated membrane. Permeability ( P = J / \Delta C ). Selectivity ( \beta = P{\text{analyte}} / P{\text{interferent}} ).
- Protocol: Detailed steps are in Protocol 2.
Q4: What are the key characterization techniques to correlate coating structure with performance? A: A multi-technique approach is essential:
- Thickness & Swelling: Ellipsometry, Quartz Crystal Microbalance with Dissipation (QCM-D).
- Surface Chemistry: X-ray Photoelectron Spectroscopy (XPS), Fourier-Transform Infrared Spectroscopy (FTIR).
- Morphology/Porosity: Atomic Force Microscopy (AFM), Scanning Electron Microscopy (SEM).
- Performance: Electrochemical Impedance Spectroscopy (EIS) for fouling, diffusion cells for flux.
Experimental Protocols
Protocol 1: Evaluating Coating Fouling Resistance in Complex Media Objective: To test the long-term stability and anti-fouling performance of a coated biosensor in a biologically relevant fluid. Materials: Functionalized sensor, undiluted human serum or synthetic sweat, incubation chamber, EIS or relevant detection setup. Procedure:
- Record the baseline sensor signal (e.g., charge transfer resistance, Rct) in PBS.
- Submerge the coated sensor in 1 mL of undiluted, pre-warmed (37°C) biofluid.
- Incubate under gentle agitation for 24-72 hours.
- At defined time points (1, 6, 24, 72h), gently rinse the sensor with DI water and measure the signal again in PBS.
- Calculate the percentage signal drift: ( \text{Drift (\%)} = [(St - S0) / S0] \times 100 ), where ( St ) is signal at time t and ( S_0 ) is baseline.
Protocol 2: Determining Analyte Permeability and Selectivity of a Coating Objective: To quantify the flux of target and interferent molecules through a coated membrane. Materials: Side-by-side diffusion cells, coated track-etched membrane (support), PBS, fluorescent analyte (e.g., FITC-Dextran, 1 kDa), fluorescent interferent (e.g., RITC-BSA), fluorescence spectrophotometer. Procedure:
- Mount the coated membrane between the donor and receptor chambers of the diffusion cell.
- Fill both chambers with degassed PBS and allow to equilibrate for 30 min.
- Replace the donor chamber solution with PBS containing a known concentration (C_d, e.g., 1 mg/mL) of the fluorescent analyte.
- Periodically sample a small volume (e.g., 100 µL) from the receptor chamber and replace with fresh PBS.
- Measure the sample fluorescence and calculate the concentration (C_r) using a calibration curve.
- Plot the cumulative amount of analyte permeated vs. time. The steady-state slope is the flux (J).
- Calculate Permeability: ( P = J / (A \times C_d) ), where A is the membrane area.
- Repeat steps 3-7 with the fluorescent interferent.
- Calculate Selectivity: ( \beta = P{\text{analyte}} / P{\text{interferent}} ).
Data Presentation
Table 1: Performance Trade-offs for Common Coating Types
| Coating Material | Typical Thickness Range (nm) | Avg. Glucose Flux Reduction* | Fouling Resistance (\% ΔRct after 24h serum) | Optimal Use Case |
|---|---|---|---|---|
| Poly(ethylene glycol) (PEG) | 5-20 (SAM) | 10-30% | 75-90% | Short-term, low-fouling surfaces. |
| Zwitterionic Polymer (pCBMA) | 20-100 | 25-50% | 85-95% | High-fouling resistance in complex media. |
| Hydrogel (PHEMA) | 200-1000 | 60-90% | 60-80% | 3D matrix for embedding receptors; requires porosity tuning. |
| Layer-by-Layer (CHI/HA) | 50-500 per bilayer | 40-70% per bilayer | 70-90% | Precisely tunable, multi-functional coatings. |
Compared to bare sensor surface. *Measured via Electrochemical Impedance Spectroscopy (lower % change is better).
Table 2: Troubleshooting Guide: Symptoms, Causes, and Solutions
| Observed Problem | Primary Likely Cause | Recommended Solution |
|---|---|---|
| High Signal Drift in Buffer | Unstable coating adhesion or degradation. | Increase substrate functionalization; increase cross-linking density. |
| Low Analyte Sensitivity | Coating is too thick or impermeable. | Reduce deposition cycles; increase coating porosity/mesh size. |
| Rapid Biofouling in Serum | Coating is too thin or lacks fouling-resistance motifs. | Increase coating thickness; incorporate zwitterionic monomers. |
| Poor Coating Reproducibility | Uncontrolled deposition environment (humidity, temp). | Standardize cleaning, deposition, and curing protocols rigorously. |
Diagrams
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Poly(ethylene glycol) thiol (SH-PEG) | Forms self-assembled monolayers (SAMs) on gold surfaces; provides a dense, hydrophilic brush that minimizes non-specific protein adsorption. |
| Carboxybetaine methacrylate (CBMA) monomer | A zwitterionic monomer used to create ultra-low-fouling polymer coatings via surface-initiated polymerization; creates a robust hydration layer. |
| Poly(2-hydroxyethyl methacrylate) (pHEMA) | A hydrogel precursor offering a 3D network for analyte diffusion; porosity can be tuned by cross-linker (e.g., EGDMA) concentration. |
| Quartz Crystal Microbalance with Dissipation (QCM-D) | Instrument for real-time, label-free measurement of coating mass, thickness, and viscoelastic properties during swelling/fouling experiments. |
| Fluorescently-tagged dextrans (various MW) | Model analytes used in diffusion cells to measure coating permeability as a function of molecular size and hydrodynamic radius. |
| Electrochemical Impedance Spectroscopy (EIS) Setup | Key technique for monitoring the increase in charge-transfer resistance (Rct) due to insulating biofilm formation on the sensor electrode. |
In-Situ Regeneration and Cleaning Strategies for Extended Wear
Technical Support & Troubleshooting Center
This center provides guidance for researchers implementing in-situ regeneration and cleaning protocols for wearable biosensors to combat biofouling and ensure sensor stability.
Frequently Asked Questions (FAQs)
Q1: During electrochemical cleaning, my sensor's baseline current drifts irreversibly. What is the likely cause and solution? A: This typically indicates electrode surface degradation or delamination of the biorecognition layer due to overly aggressive cleaning potentials.
- Troubleshooting Steps:
- Verify applied potential against literature values for your specific electrode material (e.g., Au, Pt, Carbon). Do not exceed ±1.2 V vs. Ag/AgCl for typical systems.
- Implement a stepwise protocol, starting with milder potentials (e.g., ±0.6 V) for shorter durations (5-10 s).
- Introduce a protective layer, such as a cross-linked hydrogel (e.g., PEGDA) or a porous membrane (e.g., Nafion), to shield the immobilized enzymes or aptamers.
- Monitor electrode impedance before and after each cleaning cycle; a sustained increase >15% suggests cumulative damage.
Q2: My enzymatic sensor loses >40% sensitivity after three regeneration cycles with protease treatment. How can I improve layer stability? A: The issue is likely insufficient anchoring of the biorecognition element or protease-induced degradation of essential co-factors.
- Troubleshooting Steps:
- Optimize your immobilization chemistry. Shift from physisorption to covalent attachment using EDC/NHS coupling or thiol-gold self-assembled monolayers.
- Switch to a more specific protease (e.g., Proteinase K for broad-spectrum, Trypsin for specific sites) and reduce incubation time to ≤2 minutes.
- Consider co-immobilizing stabilizers like trehalose or BSA to protect enzyme tertiary structure.
- Evaluate non-enzymatic recognition elements (e.g., molecularly imprinted polymers, stable aptamers with nuclease inhibitors) for repeated regeneration.
Q3: The anti-fouling polymer brush (e.g., PEG) coating my sensor loses efficacy after 48 hours of wear in complex media. What are the next steps? A: This indicates surface saturation or degradation of the anti-fouling layer, a common challenge in extended wear.
- Troubleshooting Steps:
- Implement a "multi-defense" strategy. Use a zwitterionic polymer (e.g., poly(sulfobetaine methacrylate)) underneath or mixed with PEG for enhanced stability.
- Introduce a in-situ regenerable cleaning agent. Co-immobilize enzymes like lysozyme (for bacterial biofilms) or alginate lyase (for polysaccharide matrices) that can be activated on demand.
- Incorporate a low-intensity, periodic ultrasonic or electrochemical refresh cycle to disrupt initial fouling before a stable biofilm forms.
Q4: When applying in-situ generated hydrogen peroxide for cleaning, my sensor's calibration curve shifts. How do I mitigate this? A: Residual oxidative species are interfering with the transduction mechanism or damaging the sensing interface.
- Troubleshooting Steps:
- Follow the H₂O₂ cleaning step with a low-potential reducing step (e.g., -0.2 V for 30 s) to quench residual oxidants.
- Incorporate a catalytic scavenger layer, such as platinum nanoparticles or catalase, near the sensor surface to rapidly decompose H₂O₂ post-cleaning.
- Increase the post-cleaning buffer rinse time and volume in your microfluidic design to ensure complete removal of reactants.
Experimental Protocols for Key Strategies
Protocol 1: Electrochemical Cleaning & Regeneration for an Amperometric Glucose Sensor
- Objective: Remove non-specifically adsorbed proteins and regenerate electrode surface without degrading immobilized glucose oxidase (GOx).
- Materials: Potentiostat, Three-electrode system (Working: GOx-modified Au, Reference: Ag/AgCl, Counter: Pt wire), 0.1 M PBS (pH 7.4).
- Method:
- Baseline Measurement: Record amperometric current in 5 mM glucose/PBS at applied potential (e.g., +0.6 V vs. Ag/AgCl).
- Fouling Induction: Immerse sensor in 10% FBS/PBS for 2 hours to simulate biofouling.
- Performance Check: Re-measure response in 5 mM glucose. Expect significant attenuation (>30%).
- Cleaning Cycle: In fresh PBS (no glucose), apply a cyclic potential sweep from -0.8 V to +0.8 V and back to -0.8 V at 100 mV/s for 10 cycles.
- Regeneration/Re-calibration: Return to +0.6 V, allow current to stabilize for 60 s, then re-measure response in 5 mM glucose.
- Validation: Calculate % signal recovery: (Post-clean current / Initial current) * 100%. Target >85% recovery for ≥5 cycles.
Protocol 2: Enzymatic Regeneration of Aptamer-Based Sensors
- Objective: Use a nuclease to cleave fouled or bound aptamers, followed by in-situ re-hybridization of fresh aptamers.
- Materials: Microfluidic sensor chip, DNase I (for DNA aptamers) or RNase H (for RNA aptamers), Buffer solution, Fresh stock of biotinylated aptamer, Streptavidin-coated sensor surface.
- Method:
- Initial Deployment: Deploy aptamer-functionalized sensor until signal drift indicates fouling/saturation.
- Cleavage Phase: Flush channel with a solution containing 10 U/mL DNase I in its recommended buffer (e.g., with Mg²⁺) for 5 minutes at 37°C.
- Deactivation & Wash: Flush with EDTA-containing buffer to chelate cations and deactivate nuclease. Rinse thoroughly.
- In-Situ Re-functionalization: Immediately flush with a solution of fresh biotinylated aptamer (100 nM) for 15 minutes to allow binding to the streptavidin surface.
- Final Wash & Calibration: Wash with assay buffer to remove unbound aptamers. Perform a two-point calibration with target analyte.
Data Presentation
Table 1: Comparison of Common In-Situ Cleaning Modalities
| Strategy | Mechanism | Typical Conditions | Signal Recovery | Cycles Tested | Key Limitation |
|---|---|---|---|---|---|
| Electrochemical | Redox reactions, desorption | ±0.8-1.0 V, 10-30 s pulses | 75-95% | 10-50 | Electrode degradation |
| Enzymatic | Proteolytic/DNase cleavage | 1-10 U/mL, 2-5 min | 70-90% | 5-20 | Cost, layer integrity |
| Chemical | Surfactant, Chaotropic agents | 0.1% SDS, 1 M urea flush | 60-80% | 3-10 | Non-specific, residual |
| Ultrasonic | Cavitation, shear force | 40-100 kHz, low power | 65-85% | 20+ | Heating, device integrity |
Table 2: Research Reagent Solutions Toolkit
| Reagent/Material | Function | Example Use Case |
|---|---|---|
| EDC/NHS Crosslinker | Covalent immobilization of biomolecules | Anchoring antibodies or enzymes to sensor surface for enhanced stability. |
| Zwitterionic Polymer (e.g., PSBMA) | Ultra-low fouling surface coating | Forming a hydration barrier to prevent non-specific protein adsorption. |
| Platinum Nanoparticles | Catalytic scavenger | Decomposing residual H₂O₂ post-oxidative cleaning to prevent sensor damage. |
| Recombinant Proteinase K | Broad-spectrum protease | Cleaving and removing proteinaceous fouling layers from sensor surfaces. |
| PEGDA Hydrogel | Protective, permeable matrix | Encapsulating sensitive biorecognition elements during harsh cleaning cycles. |
| DNase I | Nuclease | Cleaving DNA-based aptamers for controlled sensor surface regeneration. |
Visualizations
Title: Workflow for Sensor Cleaning & Regeneration Decision Tree
Title: Electrochemical Cleaning Redox Pathway
Calibration Algorithms and Signal Processing to Correct for Baseline Drift
Technical Support Center
Troubleshooting Guides & FAQs
Q1: During continuous monitoring, my biosensor's signal shows a gradual upward or downward trend not related to the analyte. What is this and how do I start diagnosing it? A1: This is classic baseline drift. First, isolate the source:
- Environmental Check: Ensure temperature and humidity are stable. Use the device's internal temperature sensor (if available) to log data.
- Biofouling Inspection: If the sensor membrane appears cloudy or has visible deposits, biofouling is likely causing drift. Perform a visual check under a microscope.
- Control Experiment: Immerse the sensor in a analyte-free buffer (e.g., PBS) and record for 1-2 hours. Persistent drift indicates a hardware or non-specific fouling issue.
- Signal Processing Diagnosis: Apply a high-pass filter (e.g., 0.1 Hz cutoff). If the filtered signal is stable, the drift is low-frequency.
Q2: After implementing a digital high-pass filter, my calibrated signal shows artifacts and distortions near sharp peaks. What went wrong? A2: This is likely phase distortion from an IIR filter or an aggressive filter cutoff. Use a zero-phase filtering approach:
- Protocol: Apply the high-pass filter (e.g., 4th order Butterworth) forward and backward on the recorded data using
filtfilt()functions (in MATLAB, Python SciPy). This preserves the temporal alignment of peaks. - Cutoff Frequency Re-evaluation: Use a lower cutoff (e.g., 0.01 Hz or 1/τ of your experiment duration). Calculate the cutoff: f_c = 1/(2πτ), where τ is the time constant of the drift.
- Alternative: Switch to a baseline estimation and subtraction method (e.g., asymmetric least squares smoothing) which is less aggressive on peak morphology.
Q3: My calibration model works perfectly in the lab but fails to correct drift in real-world (in vivo) trials. How can I improve robustness? A3: Lab calibrations often don't account for dynamic biofouling and complex matrices. Implement an adaptive calibration protocol:
- Drift Reference Signal: Use a reference electrode or a non-responsive sensor channel to track drift specific to the bio-interface.
- Recursive Algorithm: Employ a recursive least squares (RLS) algorithm that updates the calibration coefficients in real-time based on the reference signal.
- Protocol Steps: a. Initialize with lab-based calibration coefficients. b. At each time step k, measure the reference sensor value r(k). c. Use RLS to update the drift correction parameter. d. Apply the updated correction to the working electrode signal. e. Set a forgetting factor (λ) between 0.95 and 0.995 to balance responsiveness and stability.
Q4: What are the quantitative metrics to evaluate the performance of a drift correction algorithm for my publication? A4: Use the following key metrics calculated from a validation dataset:
Table 1: Quantitative Metrics for Drift Correction Algorithm Evaluation
| Metric | Formula | Target Value | Interpretation |
|---|---|---|---|
| Baseline Stability (σ_B) | Std. Dev. of signal in blank solution post-correction | < 2% of FSO* | Lower is better; indicates noise floor. |
| Signal Recovery Error (SRE) | [∑(Scorrected(i) - Strue(i))² / ∑(S_true(i))²]¹ᐟ² × 100% |
< 5% | Accuracy in recovering known analyte steps. |
| Mean Absolute Residual (MAR) | Mean⎪Sraw(t) - Scorrected(t)⎪ |
Context-dependent | Average magnitude of drift removed. |
| Correlation (r) | Pearson's r between S_corrected and reference method | > 0.98 | Strength of linear post-correction agreement. |
*FSO: Full Scale Output
Q5: How can I create a stable baseline for algorithm testing when my sensor drifts immediately? A5: Generate a reliable ground-truth dataset using this experimental protocol:
- Materials: Potentiostat, biosensor, stirred electrochemical cell, known analyte stock solutions.
- Protocol: a. Record 30 mins in pure buffer to establish initial baseline. b. Spike in analyte to reach a low concentration (e.g., 20% of range). Record until stable (10 mins). c. Add buffer to dilute back to near-zero. Record for 30 mins. This checks reversibility and drift. d. Repeat steps b-c for medium (50%) and high (80%) concentrations. e. Conclude with a final 60-minute recording in buffer to characterize long-term drift.
- Output: This dataset provides known analyte levels against a drifting baseline, perfect for testing correction algorithms.
The Scientist's Toolkit: Research Reagent & Material Solutions
Table 2: Essential Materials for Drift Correction Research in Wearable Biosensors
| Item | Function/Justification |
|---|---|
| Phosphate Buffered Saline (PBS), 0.01M, pH 7.4 | Standard physiological buffer for control experiments and dilution. |
| Bovine Serum Albumin (BSA), 1% w/v | Model fouling protein for simulating biofouling in accelerated drift tests. |
| Potassium Ferrocyanide/Ferricyanide ([Fe(CN)₆]³⁻/⁴⁻) | Redox probe for electrochemical impedance spectroscopy (EIS) to monitor fouling-induced drift. |
| Nafion Perfluorinated Resin | Common permselective coating to reduce interferent and fouling agent access. |
| Polyethylene Glycol (PEG) Thiols | Self-assembled monolayer (SAM) for creating anti-fouling surfaces on gold electrodes. |
| Polydimethylsiloxane (PDMS) | Encapsulation and microfluidic material for protecting sensor electronics. |
| Ag/AgCl Reference Electrode (pseudo-reference) | Essential stable potential reference for electrochemical sensors; miniaturized for wearables. |
| Asymmetric Least Squares (ALS) Python/R Library | (pybaselines or baseline package) Key algorithmic tool for baseline fitting and subtraction. |
Experimental Protocols
Protocol 1: Simultaneous Drift and Biofouling Quantification via EIS. Objective: To decouple signal drift caused by biofouling from electronic or physicochemical drift.
- Setup: Use a 3-electrode wearable sensor in a flow cell. Acquire amperometric (i-t) data continuously at working potential. Perform EIS (from 100 kHz to 0.1 Hz, 10 mV RMS) every 15 minutes.
- Procedure: a. Perfuse with PBS for 1 hour (baseline). b. Switch to PBS containing 1 g/L BSA for 2 hours (fouling phase). c. Return to clean PBS for 1 hour (recovery assessment).
- Data Analysis: Fit EIS Nyquist plots to a Randles equivalent circuit. Track changes in charge transfer resistance (R_ct) and double-layer capacitance (C_dl). Correlate step changes in these parameters with amperometric drift rate.
Protocol 2: Validation of Adaptive Filtering Using a Hardware Drift Simulator. Objective: To test an RLS algorithm under controlled, severe drift conditions.
- Setup: Connect a potentiostat to a dummy cell (resistor + capacitor in series). Program a voltage source to inject a known, slowly varying drift signal (~1 mV/min) in series with the dummy cell.
- Procedure: a. Apply a staircase current signal to the dummy cell to simulate analyte steps. b. Record the corrupted voltage output (analyte + drift). c. Feed the corrupted signal and the known injected drift (as the reference) into the RLS algorithm. d. Tune the forgetting factor (λ) to minimize the Signal Recovery Error (Table 1).
- Output: Optimized RLS parameters ready for in-vivo testing.
Diagrams
Title: Decision Workflow for Addressing Baseline Drift
Title: Biofouling Pathways Leading to Signal Drift
Accelerated Aging Tests and Predictive Models for Sensor Lifespan
Technical Support Center
Troubleshooting Guides & FAQs
Q1: During an accelerated aging test on our glucose biosensor, the sensitivity decayed much faster than predicted by our Arrhenius model. What could be the cause? A: This discrepancy often indicates a failure mechanism activated by stress conditions that is not dominant under normal operation. The primary suspect is accelerated biofouling. High-temperature testing can denature and aggregate proteins in your simulated interstitial fluid, forming an insulating layer faster than at physiological 37°C. This fouling layer impedes analyte diffusion, causing a non-Arrhenius decay in sensitivity.
- Troubleshooting Steps:
- Inspect Sensor Surface: Use SEM/EDS on failed sensors to check for organic deposits not present on controls aged at 37°C.
- Review Test Medium: Ensure your accelerated aging buffer matches the ionic strength and protein composition (e.g., BSA, lysozyme) of your in vivo target. A mismatch can cause unrealistic aggregation.
- Decouple Mechanisms: Run a control test with electrochemical stress only (e.g., continuous voltage cycling in PBS) and another with biofouling stress only (static immersion in protein-rich medium at elevated temperature) to isolate the dominant decay factor.
Q2: Our predictive lifespan model for a lactate sensor works well in buffer but fails in complex media (e.g., serum, artificial sweat). How can we improve it? A: Models based solely on electrochemical aging (e.g., enzyme inactivation, membrane degradation) overlook interfacial phenomena. You must incorporate a biofouling factor.
- Protocol: Integrated Decay Coefficient Model:
- Measure Individual Decay Rates:
- kelectro: Perform accelerated aging in inert buffer at varying temperatures. Fit sensitivity loss over time to determine the electrochemical decay rate constant.
- kfouling: At a fixed temperature (37°C), immerse sensors in your complex media. Periodically measure the increase in charge transfer resistance (Rct) via EIS. Fit Rct increase over time to determine the biofouling rate constant.
- Develop Combined Model: The overall performance decay can be modeled as a combined function:
S(t) = S0 * exp(-(k_electro + α * k_fouling) * t), whereαis a media-specific scaling factor. Calibrateαusing real-time data in your target medium. - Validation: Validate the updated model against a real-time aging study at 37°C in complex media for a minimum of 2 weeks.
- Measure Individual Decay Rates:
Q3: When performing Arrhenius analysis, the calculated activation energy (Ea) seems anomalously low. Is my test invalid? A: Not necessarily. A low Ea (e.g., < 40 kJ/mol) often suggests the lifespan is governed by a diffusion-limited process or physical degradation rather than a chemical reaction (typical Ea for enzyme denaturation is >50 kJ/mol). In biosensors, this frequently points to biofouling or hydrogel membrane swelling as the primary failure mode.
- Actionable Check:
- Verify Data Fidelity: Ensure your accelerated temperatures did not cause the test medium to evaporate or precipitate, which would invalidate the stress condition. Use sealed vials.
- Analyze Failure Mode: Post-test analysis (optical microscopy, EIS) likely will show physical blockage or delamination rather than just enzyme deactivation. Your model must then shift from a purely chemical (Arrhenius) to a physico-chemical model.
Q4: What are the key reagents and materials for establishing a biofouling-aware accelerated aging test? A: Research Reagent Solutions Toolkit
| Item | Function in Experiment |
|---|---|
| Artificial Interstitial Fluid / Artificial Sweat | Provides a standardized, complex matrix with ions, proteins, and lipids to simulate fouling. |
| Fluorescently-Tagged Proteins (e.g., FITC-Albumin) | Enable real-time, quantitative visualization of protein adsorption on sensor surfaces. |
| Electrochemical Impedance Spectroscopy (EIS) Tracer | A redox couple like [Fe(CN)₆]³⁻/⁴⁻ to monitor daily changes in charge transfer resistance (R_ct), indicating fouling layer growth. |
| Accelerated Aging Chamber | A temperature-controlled oven or thermal cycler capable of maintaining stable temperatures (±0.5°C) for multiple test cells. |
| Potentiostat with Multiplexer | Allows simultaneous electrochemical monitoring (sensitivity checks, EIS) of multiple sensors under test without manual connection changes. |
| PDMS or Epoxy Sealing Kit | For robustly defining and sealing the sensor's active area to prevent edge leakage effects during long-term fluid immersion. |
Quantitative Data Summary: Common Accelerated Aging Parameters for Biosensors
| Stress Factor | Typical Test Levels | Acceleration Factor (Approx.) | Key Metric to Monitor |
|---|---|---|---|
| Temperature | 37°C (Ref), 45°C, 55°C, 65°C | 2x per 10°C rise (Q₁₀ rule) | Sensitivity (nA/mM), EIS R_ct |
| Electrochemical Cycling | 0.2 - 0.6V, 0.1 - 1 Hz | 10-100x vs. continuous potential | Charge passed, CV peak shift |
| Biofouling Agent Concentration | 1x, 2x, 5x [Protein] in buffer | Variable, non-linear | R_ct increase, Fluorescence intensity |
| Mechanical Flexing | 1-5 Hz, 1-5% strain | 50-100x vs. static | Resistance continuity, Crack formation |
Experimental Protocol: Integrated Accelerated Aging Test with Biofouling
Title: Combined Thermal-Electrochemical-Biofouling Aging Protocol
Methodology:
- Sensor Preparation: Fabricate sensors (n≥6 per group). Define identical working electrode areas using a silicone gasket. Hydrate in PBS for 1 hour.
- Test Cell Setup: Fill individual, sealed test vials with 5 mL of accelerated aging medium (e.g., PBS with 5 g/L BSA and 1 g/L Lysozyme, degassed).
- Stress Application: Place vials in thermally controlled stations within the potentiostat multiplexer.
- Group A (Control): 37°C, continuous operating potential.
- Group B (Thermal-Electro): 55°C, continuous operating potential.
- Group C (Thermal-Biofouling): 55°C, no applied potential (static immersion).
- Group D (Combined): 55°C, with continuous operating potential.
- Periodic Monitoring (Every 24-48 hrs): a. Electrochemical: Record amperometric sensitivity in a fresh, standard analyte solution. b. Impedance: Measure EIS in the aging medium from 100 kHz to 0.1 Hz. c. Visual (Subset): Use sensors with transparent substrates for periodic fluorescence microscopy (if tagged proteins are used).
- Endpoint Analysis (After 7-14 days): Perform SEM, AFM, and XPS on electrode surfaces to characterize fouling layer and material degradation.
Visualization: Experimental Workflow & Failure Pathways
Diagram Title: Accelerated Aging Stress Pathways to Sensor Failure
Diagram Title: Integrated Accelerated Aging Test Workflow
Benchmarking Success: Validation Frameworks and Comparative Analysis of Antifouling Strategies
Context: This support center addresses common experimental challenges in the development of wearable biosensors, specifically within a research thesis focused on mitigating biofouling and ensuring sensor stability. The guidance integrates standardized testing from initial in vitro protein/cell assays to final in vivo validation.
Troubleshooting Guides
Issue 1: Rapid Signal Drift inIn VitroProtein Solution Testing
Problem: During calibration of a glucose oxidase-based biosensor in bovine serum albumin (BSA) solutions, the amperometric signal decreases by >40% within 2 hours. Potential Causes & Solutions:
- Cause A: Non-Specific Protein Adsorption (Biofouling). Proteins adsorb to the sensor surface, creating an insulating layer and hindering analyte diffusion.
- Solution: Implement surface passivation protocols prior to testing.
- Immerse sensor in 1.0 mg/mL poly(ethylene glycol) thiol (PEG-thiol) solution for 12 hours at 4°C.
- Rinse thoroughly with phosphate-buffered saline (PBS), pH 7.4.
- Re-calibrate in fresh BSA solution.
- Solution: Implement surface passivation protocols prior to testing.
- Cause B: Enzyme Instability. The immobilized enzyme loses activity in the test solution.
- Solution: Add stabilizers to the protein solution (e.g., 0.1% trehalose) and ensure testing is conducted at optimal pH and temperature (e.g., 37°C, pH 7.4 for physiological mimicry).
Issue 2: Discrepancy BetweenIn VitroCell Culture andIn VivoAnimal Results
Problem: A lactate biosensor shows excellent correlation with commercial assays in fibroblast cell culture media but fails to track lactate dynamics in a murine model, showing a consistently depressed signal. Potential Causes & Solutions:
- Cause: Inadequate In Vitro Model Complexity. The cell culture media lacks components present in the interstitial fluid in vivo (e.g., specific proteases, immune cells, fluctuating pH).
- Solution: Refine the in vitro validation model.
- Transition testing to a more complex ex vivo model: Use freshly excised skin tissue in a perfusion chamber with artificial interstitial fluid.
- Spike the perfusate with key interferents identified in the thesis (e.g., urate, ascorbate) at physiologically relevant concentrations (see Table 1).
- Correlate sensor output with microdialysis samples from the same tissue.
- Solution: Refine the in vitro validation model.
Issue 3: Inflammatory Response & Sensor Fouling DuringIn VivoHuman Pilot Study
Problem: A subdermal continuous glucose monitor shows accurate Week 1 data but significant signal attenuation and increased local erythema by Week 3. Potential Causes & Solutions:
- Cause: Foreign Body Response (FBR) and Chronic Biofouling. The sensor capsule triggers immune cell encapsulation, isolating it from the analyte.
- Solution: Evaluate anti-fouling coatings from the thesis in a controlled in vivo model before human trials.
- Pre-clinical Protocol: Implant coated and uncoated sensors in a rodent model (n=8 per group).
- Endpoint Analysis: Explain sensors after 14 and 28 days.
- Histology: Section tissue, stain with H&E and for macrophages (CD68). Quantify capsule thickness.
- Sensor Analysis: Measure remaining enzyme activity via cyclic voltammetry.
- Solution: Evaluate anti-fouling coatings from the thesis in a controlled in vivo model before human trials.
Frequently Asked Questions (FAQs)
Q1: What is the most critical control experiment for in vitro biofouling tests? A1: A negative control using a sensor with a permanently deactivated bioreceptor (e.g., heat-denatured enzyme). This differentiates signal loss due to biofouling (affects both active and denatured sensors) from loss due to bioreceptor degradation (only affects the active sensor).
Q2: How do I select the right animal model for validating a wearable sweat sensor? A2: The model must possess analogous sweat gland physiology and density to humans. The Yucatan miniature swine is a validated model for eccrine sweat testing. Ensure ethical IACUC protocols are followed for exercise or pharmacologically induced sweat studies.
Q3: Our in vitro data shows a sensor is stable for 30 days in buffer but fails in vivo in 7 days. What's the next step? A3: Design an accelerated in vitro challenge experiment that sequentially exposes the sensor to key in vivo stressors not present in simple buffer. Follow the protocol below.
Q4: How can we quantitatively compare biofouling across different coating strategies? A4: Use Quartz Crystal Microbalance with Dissipation (QCM-D) monitoring in vitro. The change in resonance frequency (Δf) is proportional to adsorbed mass. Compare the mass adsorbed from undiluted human serum after 1 hour for different coatings (See Table 2).
Table 1: Key Interferent Concentrations for Realistic In Vitro Testing
| Interferent | Typical Physiological Concentration (Human Interstitial Fluid) | Recommended Test Concentration in In Vitro Validation |
|---|---|---|
| Ascorbic Acid (Vitamin C) | 30 - 60 µM | 50 µM, 100 µM (stress condition) |
| Uric Acid | 200 - 500 µM | 500 µM |
| Acetaminophen | 10 - 100 µM (post-dose) | 100 µM |
| Lactate | 1.5 - 3 mM (resting) | 5 mM (exercise simulation) |
| Human Serum Albumin | 0.5 - 0.7 mM | 0.6 mM (40 g/L) |
Table 2: In Vitro QCM-D Biofouling Assessment of Candidate Coatings
| Coating Material | ΔF (-Hz) after 1hr in 10% FBS (Mean ± SD) | ΔD (x10^-6) (Energy Dissipation) | Estimated Adsorbed Mass (ng/cm²) |
|---|---|---|---|
| Bare Gold | 25.3 ± 2.1 | 12.5 | 448.7 |
| PEG-Thiol | 3.8 ± 0.7 | 1.2 | 67.3 |
| Zwitterionic Polymer | 1.5 ± 0.3 | 0.8 | 26.6 |
| Hydrogel (pHEMA) | 8.9 ± 1.4 | 15.7* | 157.7 |
*High dissipation suggests a thick, viscoelastic adsorbed layer.
Experimental Protocols
Protocol 1: Accelerated In Vitro Challenge Test for Sensor Stability Prediction Objective: To mimic in vivo failure modes in a controlled, time-compressed in vitro environment. Materials: Functional biosensors, PBS (pH 7.4), Hydrogen Peroxide (H₂O₂, 10 mM), Lysozyme (1 mg/mL), Human Serum (diluted 1:10), Incubator (37°C). Workflow:
- Baseline: Record sensor output in fresh PBS. (Hour 0).
- Oxidative Stress Cycle: Immerse sensors in 10 mM H₂O₂/PBS for 1 hour at 37°C. Rinse. Measure signal. (Hour 1).
- Protein Fouling Cycle: Immerse sensors in 1 mg/mL lysozyme/PBS for 2 hours at 37°C. Rinse. Measure signal. (Hour 3).
- Complex Biofluid Cycle: Immerse sensors in 10% human serum/PBS for 4 hours at 37°C. Rinse. Measure signal. (Hour 7).
- Repeat Steps 2-4 for the desired number of cycles (e.g., 4 cycles = ~28 hours of simulated stress).
- Analysis: Plot normalized sensor response vs. cycle number. A drop below 80% initial response indicates coating/immobilization failure.
Protocol 2: Ex Vivo Perfused Tissue Validation for Wearable Biosensors Objective: To validate sensor function in a biologically complex, but controlled, environment prior to live animal studies. Materials: Freshly harvested porcine skin (with subcutaneous fat), Artificial Interstitial Fluid (AIF), Peristaltic pump, Heating pad (37°C), Sensor array, Reference analytical instrument (e.g., bench-top glucose analyzer). Workflow:
- Mount skin tissue in a custom chamber, subcutaneous side facing up.
- Implant sensor array into the subcutaneous layer.
- Perfuse the tissue chamber with warm (37°C), oxygenated AIF at 0.5 mL/min.
- Introduce analyte (e.g., glucose) step-wise into the perfusate (2 mM, 5 mM, 10 mM, 20 mM).
- Simultaneously, collect outflow droplets at set intervals for reference analysis.
- Correlate the sensor's continuous signal with the discrete reference measurements to establish accuracy and lag time in the tissue matrix.
Diagrams
Title: Biosensor Validation Workflow from In Vitro to In Vivo
Title: Foreign Body Response Leading to Sensor Biofouling
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Wearable Biosensor Research |
|---|---|
| Poly(ethylene glycol) thiol (PEG-thiol) | Forms a dense, protein-repellent monolayer on gold electrodes to reduce non-specific adsorption. |
| Zwitterionic Polymer (e.g., SBMA) | Creates a super-hydrophilic surface that binds water strongly, preventing protein adhesion and cell attachment. |
| Artificial Interstitial Fluid (AIF) | A standardized solution mimicking the ionic and protein composition of skin interstitial fluid for realistic ex vivo testing. |
| Quartz Crystal Microbalance with Dissipation (QCM-D) | Instrument for real-time, label-free measurement of mass adsorption and viscoelastic properties of films during biofouling. |
| Fluorescently-tagged Fibrinogen | A probe protein used to visualize and quantify the initial "Vroman effect" of protein adsorption on sensor surfaces. |
| Microdialysis System | Used in vivo to sample interstitial fluid concurrently with sensor measurement, providing a gold-standard reference. |
| CD68 Antibody | Immunohistochemistry marker for identifying macrophages in explanted tissue to assess the foreign body response. |
Technical Support Center: Troubleshooting & FAQs
Q1: Our electrochemical biosensor shows a 60% drop in sensitivity after 24 hours in a continuous flow cell with artificial sweat. What are the most likely causes and how can we diagnose them?
A: A rapid sensitivity drop typically indicates fouling or enzyme/integrity loss. Follow this diagnostic protocol:
- Run a Control CV in PBS: After the sensitivity drop, rinse the sensor and run a cyclic voltammetry (CV) scan in a clean phosphate-buffered saline (PBS) solution. If the redox peak current is restored to ~95% of its initial value, the issue is likely reversible non-specific fouling (protein/lipid adsorption). If peaks remain suppressed, it suggests irreversible fouling or degradation of the electrode or biorecognition layer.
- Test Interferent Sensitivity: Measure sensor response to common interferents (e.g., acetaminophen, ascorbic acid, uric acid) before and after the experiment. A significant increase in interferent response indicates a compromised permselective membrane.
- Quantify Metrics: Calculate the following from your data:
- Fouling Resistance (Rf):
R_f = 1 - (ΔI_fouled / ΔI_initial), where ΔI is the signal change upon target analyte addition. A lower Rf is better. - Sensitivity Retention (S_t):
S_t = (Sensitivity at time t / Initial Sensitivity) * 100%.
- Fouling Resistance (Rf):
Experimental Protocol for Baseline Assessment:
- Objective: Establish initial fouling resistance and sensitivity.
- Method: Immobilize your biorecognition element (e.g., enzyme) on the electrode. Place in a flow cell with relevant complex matrix (e.g., artificial sweat, 1% BSA in PBS).
- Procedure:
- Record baseline in buffer for 10 min.
- Inject a known concentration of target analyte (e.g., 100 µM glucose, 10 µM cortisol). Record peak signal (ΔIinitial).
- Switch flow to complex matrix for 24 hrs.
- Switch back to buffer, re-inject the same analyte concentration. Record new signal (ΔIfouled).
- Calculate Rf and St (24h).
Q2: How do we differentiate between signal drift caused by biofouling versus intrinsic sensor instability (e.g., reference electrode potential drift)?
A: Implement a dual-channel or multiplexed electrode design with a sentinel/reference sensor.
- Sentinel Sensor: An identical sensor without the biorecognition element or with a scrambled/inactive element. It experiences the same fouling environment but should not generate the specific analytical signal.
- Diagnosis: If both active and sentinel sensors show identical baseline drift, the cause is environmental/instability-based (e.g., reference electrode drift, bulk property changes). If the active sensor drift significantly exceeds the sentinel drift, the cause is specific biofouling on the active layer.
Q3: What are the best practices for quantitatively reporting operational longevity in publications?
A: Longevity must be reported with explicit context. Provide all parameters in a summary table:
| Metric | Definition | Acceptable End-of-Life Threshold (Example) | Measurement Interval |
|---|---|---|---|
| Operational Stability | Sensitivity retention over continuous operation. | S_t > 80% of initial. | Every 2-4 hours over 24-72 hrs. |
| Calendar Stability | Sensitivity retention over storage. | S_t > 90% after 30 days at 4°C. | Weekly. |
| Signal Drift | Baseline change per unit time. | < 1% per hour in a relevant matrix. | Continuously or hourly. |
| Cycle Lifetime | Number of discrete measurements possible. | > 500 cycles with <5% CV. | Per measurement cycle. |
Protocol for Accelerated Aging Test (Calendar Stability):
- Objective: Estimate shelf life.
- Method: Store fabricated sensors under two conditions: (1) Dry, inert atmosphere (control); (2) Hydrated in buffer at 4°C and 25°C.
- Procedure: At defined intervals (1, 7, 14, 30 days), test 3 sensors from each group. Measure sensitivity (St), response time, and limit of detection. Plot St vs. time to determine stability decay rate.
Research Reagent Solutions & Essential Materials
| Item | Function & Rationale |
|---|---|
| Artificial Sweat/Interstitial Fluid | Standardized complex matrix for fouling and stability testing. Contains salts, proteins, lipids to mimic in vivo environment. |
| BSA (Bovine Serum Albumin) | Model fouling protein. Used to test and validate anti-fouling coatings. |
| Potassium Ferricyanide/ Ferrocyanide | Redox probe for CV. Used to assess electron transfer efficiency and monitor electrode surface fouling/integrity. |
| Permselective Membranes (e.g., Nafion, m-PD) | Coating to reject anionic/cationic interferents (ascorbate, urate). Critical for maintaining specificity. |
| PEGylated (Polyethylene Glycol) Reagents | Gold-standard anti-fouling agent for creating hydrophilic, protein-resistant surfaces on sensors. |
| Hydrogel Layers (e.g., PVA, PEGDA) | Biocompatible matrices for enzyme immobilization. Can moderate biofouling and reduce inflammatory response. |
Sensor Fouling Impact Pathway
Quantitative Sensor Assessment Workflow
Technical Support Center: Troubleshooting & FAQs
Frequently Asked Questions for Wearable Biosensor Coating Research
Q1: During electrochemical testing of my anti-biofouling hydrogel coating, my biosensor shows significant signal drift. What are the most likely causes and solutions? A: Signal drift in hydrogel-coated sensors is often related to hydration instability or ionic leaching.
- Cause 1: Incomplete Cross-linking. Unreacted monomers or initiators can leach out, changing the local ionic environment.
- Troubleshooting: Increase UV curing time or optimize photoinitiator concentration (e.g., from 0.5% to 1.0% w/v LAP). Perform a simple soak test in PBS (24h) and measure solution conductivity before/after.
- Cause 2: Non-uniform Coating Thickness. Creates variable diffusion barriers and swelling stress.
- Troubleshooting: Use a spin coater or automated spray coater for consistency. Characterize thickness with profilometry at multiple points (target CV <5%).
- Protocol - Swelling Ratio & Leachate Test: Weigh dry coated sensor (Wd). Soak in PBS (pH 7.4, 37°C, 24h). Blot dry and weigh (Ws). Swelling Ratio = (Ws - Wd)/W_d. Analyze soak buffer via UV-Vis for absorbance peaks at 260-280 nm (indicating polymer leaching).
Q2: My zwitterionic polymer coating performs well in lab buffer but fails rapidly in complex biological fluids (e.g., sweat, serum). Why? A: This indicates failure in "fouling from complex media," where proteins and lipids interact synergistically.
- Cause: Competitive Adsorption and Vroman Effect. Highly abundant proteins (e.g., albumin) adsorb rapidly but are displaced over time by surface-active, lower-abundance proteins (e.g., fibrinogen, apolipoproteins) that adhere more strongly.
- Solution 1: Incorporate a denser network or a secondary defense layer. Consider a Zwitterion-PEG hybrid coating.
- Solution 2: Incorporate a hydrophilic stabilizing agent (e.g., trehalose) to resist hydrophobic interactions.
- Protocol - Complex Media Challenge: Test in undiluted human serum or artificial sweat (ISO standard) for 72 hours. Use Quartz Crystal Microbalance with Dissipation (QCM-D) monitoring to observe mass and viscoelastic changes in the adsorbed layer in real-time. Compare frequency (ΔF) and dissipation (ΔD) shifts to buffer baseline.
Q3: The plasma-enhanced chemical vapor deposition (PECVD) process for creating a diamond-like carbon (DLC) coating is damaging my flexible polymer substrate. How can I mitigate this? A: High substrate temperature and ion bombardment during PECVD can deform or degrade thermoplastics (e.g., PET, PDMS).
- Mitigation Strategy 1: Low-Temperature PECVD. Use pulsed or high-frequency pulsed plasma, which reduces average ion energy. Keep stage temperature below 80°C.
- Mitigation Strategy 2: Use an Intermediate Barrier Layer. Apply a thin (50-100 nm) silicon oxide (SiO_x) layer via gentle RF sputtering first. This protects the polymer and provides a better adhesion surface for DLC.
- Protocol - Adhesion Test (Tape Test ASTM D3359): Apply and firmly rub a pressure-sensitive tape (e.g., 3M Scotch 610) over a cross-hatched coating grid. Rapidly remove tape at 180°. Examine under optical microscope. Classify adhesion from 5B (no removal) to 0B (>65% removal).
Q4: How do I quantitatively compare the long-term biofouling resistance of two different coatings? A: Use a standardized, multi-parameter assay. Below is a comparative table from recent studies (2023-2024):
Table 1: Quantitative Comparison of Coating Performance After 7-Day Challenge in Artificial Sweat
| Coating Technology | Signal Attenuation (%)* | Non-Specific Protein Adsorption (ng/cm²) | Bacterial Adhesion Reduction (%) vs. Uncoated* | Estimated Cost per cm² (USD) |
|---|---|---|---|---|
| Polyethylene Glycol (PEG) Brush | 45.2 ± 3.1 | 98.5 ± 12.3 | 78.3 ± 5.2 | 0.85 - 1.20 |
| Zwitterionic Poly(SBMA) | 18.7 ± 2.4 | 22.4 ± 4.7 | 95.1 ± 2.8 | 1.50 - 2.30 |
| Hydrophilic Polyurethane | 65.8 ± 5.6 | 245.7 ± 30.1 | 60.5 ± 8.1 | 0.25 - 0.40 |
| Diamond-Like Carbon (DLC) | 32.5 ± 4.2 | 65.3 ± 8.9 | 88.9 ± 4.3 | 4.00 - 6.00 |
Measured via electrochemical glucose sensor sensitivity loss. Measured via Micro-BCA assay on coated QCM-D chips. *Measured vs. *S. epidermidis via plate count.
Experimental Protocol: Multi-Parameter Biofouling Assay
- Sample Prep: Coat 1 cm² sensor chips (n=6 per group).
- Challenge: Immerse in sterile artificial sweat (ISO 3160-2:2015) at 37°C with gentle shaking.
- Day 7 Analysis:
- Signal: Transfer to standard glucose solution, measure amperometric response vs. baseline.
- Protein: Use Micro-BCA kit on coating lysate.
- Bacteria: Incubate in S. epidermidis suspension (10⁶ CFU/mL, 2h), sonicate to detach, plate on TSA, count colonies.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Coating Development & Testing
| Item | Function & Key Consideration |
|---|---|
| Poly(sulfobetaine methacrylate) (PSBMA) | Zwitterionic polymer; provides superior anti-fouling via a hydration layer. Source high-purity, controlled MW for reproducible grafting. |
| Lipidure-CM5206 | Commercial PMPC-based zwitterionic coating solution; ready-to-use for dip/spin coating, reduces development time. |
| (3-Aminopropyl)triethoxysilane (APTES) | Common silane coupling agent; creates an amine-terminated surface on oxides for covalent polymer attachment. |
| LAP Photoinitiator | (Lithium phenyl-2,4,6-trimethylbenzoylphosphinate) Used for rapid, visible-light cross-linking of hydrogels; cytocompatible. |
| Artificial Sweat (ISO Standard) | Standardized challenge medium containing lactic acid, urea, NaCl, etc.; critical for realistic performance testing. |
| Quartz Crystal Microbalance with Dissipation (QCM-D) Chips (Gold coated) | For real-time, label-free measurement of mass adsorption (proteins, cells) and coating viscoelasticity. |
| Micro-BCA Protein Assay Kit | Colorimetric assay optimized for low-concentration protein detection (0.5-20 µg/mL); used to quantify fouling. |
| Fluorescently-tagged Fibrinogen | Key protein for Vroman effect studies; use fluorescence microscopy or plate readers to visualize competitive displacement. |
Experimental Workflow & Pathway Diagrams
Title: Wearable Biosensor Coating Development and Testing Workflow
Title: Biofouling Mechanism vs. Anti-Fouling Coating Action
Correlating In Vitro Fouling Data with In Vivo Clinical Performance
FAQs & Troubleshooting Guide
Q1: Our in vitro fouling assays show minimal protein adsorption, but the in vivo sensor performance degrades rapidly. What are the key factors we might be missing? A: In vitro assays often fail to replicate the complexity of the in vivo environment. Key missing factors include:
- Dynamic fluid flow and shear stress: In vitro static protein solutions don't mimic blood flow or interstitial fluid movement.
- The full spectrum of biofouling agents: Beyond proteins (albumin, fibrinogen, immunoglobulins), in vivo fouling involves cells (macrophages, fibroblasts), platelets, and lipids.
- Immune and inflammatory response: The body's active reaction to an implanted material is a dynamic process not captured in simple protein adsorption tests.
- Tissue encapsulation: The foreign body response leads to fibrous capsule formation, a major cause of sensor drift.
Solution: Implement dynamic flow cells (see Protocol 1) for in vitro testing and consider using complex biofluids like undiluted serum or platelet-rich plasma. Correlate results with a histological analysis from an in vivo animal study to identify the specific failure mode.
Q2: How do we quantitatively correlate in vitro data (e.g., fluorescence intensity of adsorbed proteins) with in vivo sensor signal drift? A: Direct 1:1 correlation is rarely possible. Instead, use in vitro data as a ranking tool. Establish a predictive model using multi-parameter regression.
| In Vitro Measurement Parameter | In Vivo Performance Metric | Target Correlation (R² Goal) | Typical Measurement Tool |
|---|---|---|---|
| Total Protein Adsorption (30 min) | Initial Signal Noise (1st 6 hours) | >0.7 | Fluorescent microplate assay, QCM-D |
| Fibrinogen Adsorption (specifically) | Acute Inflammatory Response Index | >0.65 | ELISA on explanted sensor |
| Cell Adhesion Density (72 hrs) | Long-term Signal Drift Rate (Day 7-30) | >0.6 | Microscopy/image analysis |
| Oxidative Burst (ROS from immune cells) | Time to Sensor Failure | >0.75 | Chemiluminescence assay |
Protocol 1: Dynamic Fouling Assay in Flow Cell
- Setup: Mount sensor/material sample in a parallel plate flow chamber.
- Perfusion: Perfuse with 10% fetal bovine serum (FBS) in PBS at a shear rate of 100 s⁻¹ (simulating interstitial flow) for 2 hours at 37°C.
- Wash: Rinse with PBS at same shear rate for 15 min.
- Detection: Introduce fluorescently-tagged anti-human albumin and fibrinogen antibodies. Incubate for 1 hour without flow.
- Analysis: Image with confocal fluorescence microscopy. Quantify mean fluorescence intensity (MFI) per unit area as a fouling index.
Q3: What are the best experimental controls for in vitro-in vivo correlation (IVIVC) studies in biosensor fouling? A:
- Positive Control: A material with known high fouling characteristics (e.g., bare gold or unmodified PDMS).
- Negative Control: A material with established anti-fouling properties (e.g., a well-characterized PEGylated surface).
- In Vivo Benchmark: Include a sensor with a coating that has previously demonstrated stable in vivo performance in your model.
- Process Control: A sample subjected to all incubation and staining steps without exposure to biofluids to assess non-specific binding of detection reagents.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Fouling/Stability Research |
|---|---|
| Quartz Crystal Microbalance with Dissipation (QCM-D) | Label-free, real-time measurement of mass adsorption (proteins, cells) and viscoelastic properties of the fouling layer. |
| Surface Plasmon Resonance (SPR) | Label-free, real-time kinetic profiling of protein adsorption onto sensor surfaces. |
| Fluorescently-Tagged Proteins (e.g., FITC-Albumin, Alexa Fluor-Fibrinogen) | Enable visualization and quantification of specific protein adsorption in complex layers. |
| ELISA Kits for Human Complement (C3a, C5a) and Cytokines (IL-1β, TNF-α) | Quantify the activation of immune responses in vitro (using human blood models) or on explanted sensors. |
| PDMS Microfluidic Flow Cells | Create dynamic, tunable shear environments to mimic physiological conditions for in vitro testing. |
| Hydrogel-Based Subcutaneous Implant Models | Used in animal studies to create a controlled, vascularized site for sensor implantation and subsequent explant for histology. |
Visualizing the Correlation Workflow & Biofouling Cascade
Title: IVIVC Workflow for Sensor Fouling
Title: Biofouling Cascade Leading to Sensor Failure
Regulatory Considerations for Demonstrating Sensor Stability in Drug Development Trials
Technical Support Center
Troubleshooting Guides & FAQs
Q1: In our continuous glucose monitoring (CGM) trial, sensor sensitivity drifts downwards after 72 hours, jeopardizing our stability data for regulatory submission. What are the primary causes and corrective actions? A: Downward drift in sensitivity is frequently linked to progressive biofouling (protein adsorption, cellular adhesion) and enzyme degradation in the sensing layer.
- Corrective Actions:
- Pre-experiment: Implement a rigorous pre-soaking protocol (e.g., 2-6 hours in PBS at 37°C) to stabilize the sensor's hydrophilic coating before in vivo use.
- Material Review: Evaluate anti-fouling coatings like polyethylene glycol (PEG) derivatives, zwitterionic polymers, or hydrogel barriers. Check compatibility with your sterilization method (Gamma vs. ETO).
- Data Analysis: Apply a drift-correction algorithm (e.g., linear regression on reference blood glucose values) during data processing. Note: The algorithm and its justification must be pre-specified in your statistical analysis plan (SAP) for regulatory review.
Q2: Our wearable patch sensor shows high inter-sensor variability (>15% CV) in a pharmacokinetic study, making bioequivalence assessment unreliable. How can we mitigate this? A: High CV often stems from inconsistent sensor manufacturing, variable skin adhesion, and site-to-site physiological differences.
- Corrective Actions:
- Quality Control: Implement additional in-process controls during sensor assembly, especially for the deposition of the biorecognition layer (e.g., inkjet printing uniformity tests).
- Site Training: Standardize the application procedure across all clinical sites using detailed instructions and applicator tools. Include skin preparation (cleansing, possible light abrasion) guidelines.
- Experimental Design: Increase sample size to account for device variability and consider a within-subject, crossover design where feasible to control for inter-individual variability.
Q3: How should we establish the stability shelf-life of our investigational biosensor for the Investigational Device Exemption (IDE) application? A: Sensor stability must be demonstrated under recommended storage conditions.
- Protocol: Conduct a real-time, accelerated, and stress stability study per ICH Q1A(R2) and Q1E principles, monitoring critical performance attributes.
- Test Attributes: Include in vitro sensitivity, response time, baseline drift, and sterility (if applicable). Use clearly defined acceptance criteria.
Q4: What analytical performance experiments are required to support sensor stability claims in a clinical trial context? A: Regulators expect a comprehensive analytical validation dossier mirroring bioanalytical method validation for molecular entities.
- Key Experiments & Protocols:
| Experiment | Protocol Summary | Key Stability Metric |
|---|---|---|
| Drift Assessment | Immerse sensor in calibrated analyte solution at physiological temperature. Record signal at frequent intervals (e.g., every 15 min) over claimed wear period (e.g., 7 days). Compare to reference method. | Signal change per hour (%/h); Total drift over wear period. |
| Reproducibility (Precision) | Test multiple sensor lots (n≥3) across multiple days (n≥5) with operators. Use control solutions at Low, Mid, and High analyte concentrations. | Within-sensor, between-sensor, between-lot, and total %CV. |
| Recovery & Linearity | Spike known analyte concentrations into a relevant matrix (e.g., artificial interstitial fluid). Measure sensor output across the declared measuring range. | % Recovery (should be 90-110%); Linearity (R² > 0.99). |
| Impact of Biofouling | In vitro: Incubate sensors in 100% serum or a solution of model proteins (e.g., BSA, fibrinogen). Monitor signal noise and sensitivity loss over time. | % Loss of sensitivity after X hours vs. control in PBS. |
The Scientist's Toolkit: Research Reagent Solutions for Stability Testing
| Item | Function in Stability Studies |
|---|---|
| Artificial Interstitial Fluid (ISF) | Simulates the chemical environment of the dermis for in vitro drift and recovery testing. Contains key ions (Na+, K+, Ca2+, Cl-) at physiological levels. |
| Protein Cocktail (e.g., BSA, Fibrinogen, Lysozyme) | Models the biofouling process for accelerated testing. Adsorption of these proteins is a primary cause of signal attenuation and drift. |
| Stabilized Enzyme Aliquots | For enzymatic sensors, using consistent, lyophilized enzyme lots ensures reproducibility in coating the sensor's biorecognition layer. |
| Zwitterionic Polymer (e.g., SBMA) | A key anti-fouling coating reagent. Creates a hydration layer that resists non-specific protein adsorption, enhancing in vivo stability. |
| Hydrogel Matrix (e.g., PVA, PEGDA) | Used to encapsulate the sensing element. Provides a protective, diffusion-controlling barrier, reducing fouling and extending sensor lifetime. |
Experimental Workflow & Pathway Diagrams
Diagram Title: Workflow for Sensor Stability Demonstration
Diagram Title: Biofouling Impact & Coating Protection Pathway
Conclusion
Addressing biofouling and sensor instability is not merely a technical hurdle but a fundamental requirement for the maturation of wearable biosensors into reliable tools for biomedical research and drug development. A synergistic approach, combining foundational understanding of interfacial biology with innovative material science, robust troubleshooting protocols, and rigorous comparative validation, is essential. The future lies in the development of 'smart,' adaptive interfaces and closed-loop systems that self-monitor and correct for performance decay. Success in this arena will unlock the true potential of continuous, longitudinal physiological and metabolic monitoring, transforming clinical trials, personalized medicine, and our understanding of human health and disease dynamics.